 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA03g35060 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92059.4 Da PI: 6.4662 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.9 | 8.1e-19 | 15 | 73 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
CA03g35060 15 KYVRYTPEQVEALERLYHDCPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 73
5679*****************************************************97 PP
| |||||||
| 2 | START | 170.5 | 1.1e-53 | 160 | 367 | 3 | 205 |
HHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEEEXXT CS
START 3 aeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmvaelq 99
aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++ +++++v+ ++ g+++l +++l+
CA03g35060 160 AEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCTGVAARACGLVGLEPT-RVSEILKDRPSWFRDCRTVDVINVLPTAngGTIELLYMQLY 257
7899*****************************************************.9999999999****************9999*********** PP
TXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHH CS
START 100 alsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglaegak 194
a+++l+p Rdf+ +Ry+ + +g++v++++S+ ++q+ p+ +++vRae lpSg+li+p+++g+s +++v+h++l+++ ++++lr+l++s+++ ++k
CA03g35060 258 APTTLAPaRDFWLLRYTTVMDDGSLVVCERSLGNTQNGPSmppVPNFVRAEILPSGYLIRPCEGGGSIIHIVDHMNLEAWCVPEVLRPLYESSAVLAQK 356
*************************************9999999******************************************************* PP
HHHHHTXXXXX CS
START 195 twvatlqrqce 205
t+va+l+++++
CA03g35060 357 TTVAALRYLRQ 367
******99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.516 | 10 | 74 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.2E-16 | 12 | 78 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 9.41E-17 | 14 | 77 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 7.98E-17 | 15 | 75 | No hit | No description |
| Pfam | PF00046 | 2.2E-16 | 16 | 73 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-18 | 17 | 73 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.77E-6 | 67 | 106 | No hit | No description |
| PROSITE profile | PS50848 | 25.314 | 149 | 377 | IPR002913 | START domain |
| CDD | cd08875 | 4.59E-75 | 153 | 369 | No hit | No description |
| SMART | SM00234 | 1.4E-41 | 158 | 368 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.48E-38 | 159 | 370 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 1.3E-22 | 159 | 363 | IPR023393 | START-like domain |
| Pfam | PF01852 | 1.3E-51 | 160 | 367 | IPR002913 | START domain |
| Pfam | PF08670 | 1.1E-49 | 694 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MASCKDGKSV MDNGKYVRYT PEQVEALERL YHDCPKPSSM RRQQLIRECP ILSNIEPKQI 60 KVWFQNRRCR EKQRKEASRL QTVNRKLTAM NKLLMEENDR LQKQVSQLVY ENGYFRRQTH 120 STPLATKDTS CESVVTSGQH HLTSQHPPRD ASPAGLLSTA EETLTEFLSK ATGTAVEWVQ 180 MPGMKPGPDS IGIIAISHGC TGVAARACGL VGLEPTRVSE ILKDRPSWFR DCRTVDVINV 240 LPTANGGTIE LLYMQLYAPT TLAPARDFWL LRYTTVMDDG SLVVCERSLG NTQNGPSMPP 300 VPNFVRAEIL PSGYLIRPCE GGGSIIHIVD HMNLEAWCVP EVLRPLYESS AVLAQKTTVA 360 ALRYLRQIAQ EVSQTNVTNW GRRPAALRAL GQRLSRGFNE ALNGLADEGW SMLDSDGMDD 420 VAILVNSSPD KLMGLNLSFA NGFSSMSNAV MCAKASMLLQ NVPPAILLRF LREHRSEWAD 480 NNIDAYAAAA IKVGPCSLPG ARVGNFGGQV ILPLAHTVEH EEASLLLEVI KLEGHSPEDA 540 IMPRDMFLLQ LCSGMDENAV GTCAELVFAP IDASFADDAP LLPSGFRIIS LESGKEASSP 600 NRTLDLTSAL ETGPAENKVA NDLHTVGGTS RSVMTIAFQF AFESHMQESV ASMARQYVRS 660 IISSVQRVAL ALSPSHLGSH GGLRSPLGTP EAHTLARWIC QSYRCFLGVE LLKANTDQGS 720 ESILKGLWHH SDAIICCSAK ALPVFTFANQ AGLDMLETTL VALQDITLEK IFDDHGKKNL 780 CTEFPQIMQQ GFACLQGGIC LSSMSRPISY ERAVAWKVMN EEDTAHCICF MFVNWSFV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
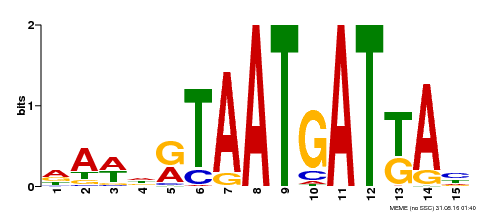
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JQ686925 | 0.0 | JQ686925.1 Nicotiana tabacum cultivar SR1 homeobox-leucine zipper protein ATHB-15-like protein (HB15) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016562656.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2G3A3Q2 | 0.0 | A0A2G3A3Q2_CAPAN; Homeobox-leucine zipper protein HOX32 | ||||
| STRING | PGSC0003DMT400006735 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA457 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



