 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA01g22800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 984aa MW: 109192 Da PI: 6.0111 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.9 | 9e-41 | 126 | 203 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C +dls+ak+yhrrhkvCe+hska+ +lv +++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+q+
CA01g22800 126 VCQVDDCGTDLSKAKDYHRRHKVCEMHSKASRALVGNVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNKRRRKTQS 203
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 8.5E-33 | 122 | 188 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.086 | 124 | 201 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 7.85E-38 | 125 | 206 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.6E-30 | 127 | 200 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 2.35E-7 | 753 | 867 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.2E-5 | 756 | 870 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.09E-6 | 757 | 867 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 984 aa Download sequence Send to blast |
MGGPDLRGLG KRSLEWDLTD WKWDGDLFIA TPLHQNAPNY QSRQFFPVET GNLASSNSSS 60 SCSEEVNHGM EQQRRELEKR RRVIVVEEDD SGPLSQLPHP PXLGGNWDGA AGKRTKLVAP 120 AGSRAVCQVD DCGTDLSKAK DYHRRHKVCE MHSKASRALV GNVMQRFCQQ CSRFHALQEF 180 DEGKRSCRRR LAGHNKRRRK TQSESVANNN SVNEGQASGY SLMSLLKILS NMHSNGTNPT 240 EDQNLLSHLL RSLASQGTTN GDKSLSGLLQ ESSNLLNNRS ILRNPEVASL ISNGSQAPPR 300 AKEHQFTNSA TEMPQKRLDV HDVRLEDART ASSQSPGILF PIQSNSQAYT PGRESTTGRS 360 KLMDFDLNDA YVDSDDCGED LVRSPVPECP SWLQQDSHQS SPPQTSGNSD SASAQSPSSS 420 SGDNQNRTDR IVFKLFGKDP SDFPFVVRAQ ILDWLSHSPT EIESYIRPGC IVLTIYLRLP 480 ESAWEELSYN LSSSLSRLLD VPGGDSFWTR GWIYIRVQNQ IAFVCDGQVL LDMSLPYVSN 540 DDSTLLSVRP IAVPVSGRVQ FLVKGYNLTK PSTRLLCALD GSYLVPEANN EVEEHGDVVG 600 KDDKIQSLNF TYSVSAVAGR GFIEVEDHGV NNSFFPFIIA EEDVCSEIRM LESDLELSSS 660 DYVNGQTSNM EARNQAMDFI HELGWLLHRN NLKARLDHFG PNAVLYPLKR FKWLVDFSVD 720 HDWCAVVKKL LNILLDGTVG AGDNSFLKFA LSELGLLHRA VRRNSRPLVE LLLTYTPESV 780 ADELCSEYQS LVGIDGEFLF RPDCVGPAGL TPLHVAACID GYEDVLDALT DDPGKVAIEA 840 WKNTRDGAGF TPEDYARLRG HYSYIHLVQR KISKKATSGH IVVDIPSVPA VVESNNQKDE 900 VYTTTSLEIT MTERTPIARP CGLCDRKLAY GSRGRSLLYR PAMFSMVAMA AVCVCVALLF 960 RGSPEVLYIF RPFRWEMVDF GTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-31 | 126 | 200 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 78 | 82 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
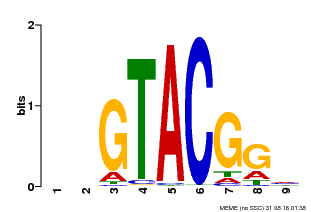
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975440 | 0.0 | HG975440.1 Solanum pennellii chromosome ch01, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016539898.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A1U8DXZ3 | 0.0 | A0A1U8DXZ3_CAPAN; squamosa promoter-binding-like protein 1 | ||||
| STRING | PGSC0003DMT400013317 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



