 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA01g19070 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 836aa MW: 91074.1 Da PI: 6.1456 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.9e-21 | 132 | 187 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
CA01g19070 132 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 187
688999***********************************************999 PP
| |||||||
| 2 | START | 202.6 | 1.7e-63 | 339 | 563 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEEC CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevis 86
ela++a++elvk+a+ +ep+W++s e +n++e++++f++ + + +ea+r++g+v+ ++ lve+l+d++ +W e+++ + +t++vis
CA01g19070 339 ELALAAMDELVKMAQTDEPLWFRSMegarEVLNQEEYMRTFTPCIGmrpntFVSEASRETGMVIINSLALVETLMDSN-KWAEMFPcliaRTSTTDVIS 436
5899*********************9****************9999********************************.******************** PP
TT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXH CS
START 87 sg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlph 178
sg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + + +++lpSg+++++++ng+skvtwveh+++++ h
CA01g19070 437 SGmggtrnGALQLMHAELQVLSPLVPiREVNFLRFCKQHAEGVWAVVDVSIDTIRETSAAPTFQSCRRLPSGCVVQDMPNGYSKVTWVEHAEYDEGANH 535
********************************************************9999*************************************** PP
HHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 179 wllrslvksglaegaktwvatlqrqcek 206
l+r+l++ g+ +ga++wvatlqrqce+
CA01g19070 536 HLYRQLISAGMGFGAQRWVATLQRQCEC 563
**************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.05E-20 | 119 | 189 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-21 | 119 | 189 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.248 | 129 | 189 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-17 | 130 | 193 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.92E-18 | 131 | 189 | No hit | No description |
| Pfam | PF00046 | 2.0E-18 | 132 | 187 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 164 | 187 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.544 | 330 | 566 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.47E-31 | 332 | 563 | No hit | No description |
| CDD | cd08875 | 7.02E-124 | 334 | 562 | No hit | No description |
| SMART | SM00234 | 7.9E-47 | 339 | 563 | IPR002913 | START domain |
| Pfam | PF01852 | 3.5E-55 | 339 | 563 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.07E-24 | 582 | 762 | No hit | No description |
| SuperFamily | SSF55961 | 9.07E-24 | 789 | 820 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 836 aa Download sequence Send to blast |
MNFGGFLDNN SGGGGARIVA DTPFNNNSSS GNNKSSNNNN NDNNMPTGAI SQPRLLPQSL 60 AKTMFNSPGL SLALQTGMEG QNDVTRMGEA YEGNNSVGRR SREEEPDSRS GSDNLEGASG 120 DEQDAADKPP RKKRYHRHTP QQIQELESLF KECPHPDEKQ RLELSKRLCL ETRQVKFWFQ 180 NRRTQMKTQL ERHENSILRQ ENDKLRAENM SIREAMRNPI CTNCGGPAMI GEISLEEQHL 240 RIENARLKDE LDRVCALAGK FLGRPISSLV TSMPPPMPNS SLELGVGSNG FGGLSNVPTS 300 LPLAPPDFGV GISASMPVVP STRQTSGIER SLERSMYLEL ALAAMDELVK MAQTDEPLWF 360 RSMEGAREVL NQEEYMRTFT PCIGMRPNTF VSEASRETGM VIINSLALVE TLMDSNKWAE 420 MFPCLIARTS TTDVISSGMG GTRNGALQLM HAELQVLSPL VPIREVNFLR FCKQHAEGVW 480 AVVDVSIDTI RETSAAPTFQ SCRRLPSGCV VQDMPNGYSK VTWVEHAEYD EGANHHLYRQ 540 LISAGMGFGA QRWVATLQRQ CECLAILMSS TVTSRDYTAI TPSGRRSMLK LAQRMTNNFC 600 AGVCASTVHK WNKLCAGNVD EDVRVMTRKS VDDPGEPPGI VLSAATSVWL PVSPQRLFDF 660 LRDERLRSEW DILSNGGPMQ EMAHIAKGQD HGNCVSLLRA SAMNANQSSM LILQETCIDA 720 AGALVVYAPV DIPAMHVVMN GGDSAYVALL PSGFSIVPDG PGSRGSSTGN CGSNGPPSCN 780 GGPDQRISGS LLTVAFQILV NSLPTAKLTV ESVETVNNLI SCTVQKIKGA LHCES* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
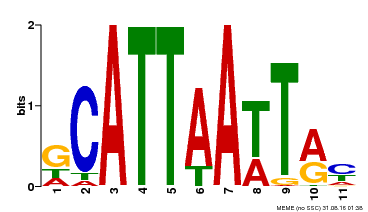
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GQ222185 | 0.0 | GQ222185.1 Solanum lycopersicum cultivar M82 cutin deficient 2 (CD2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016575742.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2G3AGV6 | 0.0 | A0A2G3AGV6_CAPAN; Homeobox-leucine zipper protein ROC6 | ||||
| STRING | PGSC0003DMT400071377 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



