 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv9_223840_krez.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 253aa MW: 28617.1 Da PI: 6.1119 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 63.6 | 3.9e-20 | 7 | 52 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEdell ++v+++G ++W+ I++ ++ gR++k+c++rw +
Bv9_223840_krez.t1 7 KGPWSPEEDELLQKLVQKHGARNWSVISKSIP-GRSGKSCRLRWCNQ 52
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 61.3 | 2e-19 | 61 | 103 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+++teEde++v+a++q+G++ W+tI+r ++ gRt++ +k++w++
Bv9_223840_krez.t1 61 PFSTEEDEIIVRAHAQHGNK-WATISRLLN-GRTDNAIKNHWNST 103
89******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 25.821 | 2 | 57 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.82E-32 | 4 | 100 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.1E-17 | 6 | 55 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.5E-20 | 7 | 52 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-26 | 8 | 60 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.66E-16 | 9 | 51 | No hit | No description |
| SMART | SM00717 | 2.0E-15 | 58 | 106 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.346 | 59 | 108 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.3E-15 | 61 | 103 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-23 | 61 | 107 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.01E-11 | 61 | 104 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:2000022 | Biological Process | regulation of jasmonic acid mediated signaling pathway | ||||
| GO:2000031 | Biological Process | regulation of salicylic acid mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 253 aa Download sequence Send to blast |
MAADRIKGPW SPEEDELLQK LVQKHGARNW SVISKSIPGR SGKSCRLRWC NQLSPEVEHR 60 PFSTEEDEII VRAHAQHGNK WATISRLLNG RTDNAIKNHW NSTLKRKFSS IVAGDEIYRP 120 EKRPEISFSP PLSPSESDVS DSGVGYLPTV VEERENQEND PLTALTLSLP GIIEEHPSKS 180 ESTQEETRKV DVVGCESVKE TMSFGPELMA VMQEMIKKEV RNYMMEAERS GVSFDEYSAR 240 NAVVNRFCNG HIH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 2e-39 | 6 | 108 | 3 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| 1h88_C | 1e-38 | 6 | 108 | 57 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 1e-38 | 6 | 108 | 57 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 2e-39 | 6 | 108 | 3 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 2e-39 | 6 | 108 | 3 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor (PubMed:23067202, PubMed:23603962). Represses the expression of protein phosphatases 2C in response to abscisic acid (ABA). Confers resistance to abiotic stresses dependent of ABA (PubMed:18162593, PubMed:9678577). In response to auxin, activates the transcription of the auxin-responsive gene IAA19. The IAA19 transcription activation by MYB44 is enhanced by direct interaction between MYB44 and PYL8 (PubMed:24894996). Transcriptional activator of WRKY70 by direct binding to its promoter region, especially at 5'-TAACNG-3' and 5'-CNGTTA-3' symmetric motifs (PubMed:23067202, PubMed:23603962). Activates salicylic acid (SA)- mediated defenses and subsequent resistance to biotrophic pathogen P.syringae pv. tomato DC3000, but represses jasmonic acid (JA)-mediated defenses responses against the necrotrophic pathogen A.brassicicola (PubMed:23067202). {ECO:0000269|PubMed:18162593, ECO:0000269|PubMed:23067202, ECO:0000269|PubMed:23603962, ECO:0000269|PubMed:24894996, ECO:0000269|PubMed:9678577}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00596 | DAP | Transfer from AT5G67300 | Download |
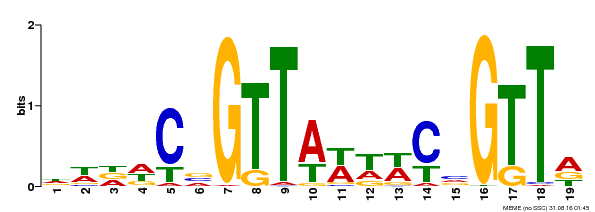
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By drought, cold, high salinity, cadmium (CdCl(2)), salicylic acid (SA), jasmonate (JA), ethylene, gibberellic acid (GA), and ABA (PubMed:16463103, PubMed:18162593, PubMed:23067202). The induction by JA is COI1-dependent (PubMed:23067202). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:18162593, ECO:0000269|PubMed:23067202}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010691674.1 | 0.0 | PREDICTED: transcription factor MYB44-like | ||||
| Swissprot | Q9FDW1 | 4e-76 | MYB44_ARATH; Transcription factor MYB44 | ||||
| TrEMBL | A0A0K9R8X8 | 1e-146 | A0A0K9R8X8_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010691674.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G67300.1 | 2e-78 | myb domain protein r1 | ||||



