 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv9_215020_hpmh.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1005aa MW: 112084 Da PI: 6.025 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 182.6 | 4.4e-57 | 21 | 136 | 3 | 118 |
CG-1 3 kekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenpt 93
+ ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+e+n+
Bv9_215020_hpmh.t1 21 EAQHRWLRPAEICEILRNHQKFQISSEPPNRPSSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEDNEY 111
349**************************************************************************************** PP
CG-1 94 fqrrcywlLeeelekivlvhylevk 118
fqrr+ywlLeee+ +iv+vhylevk
Bv9_215020_hpmh.t1 112 FQRRSYWLLEEEFMHIVFVHYLEVK 136
**********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 84.67 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.9E-82 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.2E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.5E-7 | 428 | 515 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 3.1E-6 | 430 | 515 | IPR002909 | IPT domain |
| CDD | cd00102 | 2.30E-4 | 430 | 516 | No hit | No description |
| SuperFamily | SSF81296 | 2.42E-18 | 430 | 515 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.95E-16 | 464 | 707 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.13E-12 | 584 | 703 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 4.8E-16 | 587 | 708 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 14.716 | 602 | 715 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF52540 | 4.51E-8 | 817 | 870 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.3 | 819 | 841 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.876 | 820 | 849 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0017 | 821 | 840 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0012 | 842 | 864 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 843 | 867 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 6.5E-5 | 846 | 864 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1005 aa Download sequence Send to blast |
MADRGSLNPG IRLDIQQLQL EAQHRWLRPA EICEILRNHQ KFQISSEPPN RPSSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEDNE YFQRRSYWLL 120 EEEFMHIVFV HYLEVKGNRS SIRDTEPVLS NSQISNSRSS SVSNSYNEAF TGNMGSPSAV 180 SSLTSSYEDI ESEDNHQAIS KYPSPLNFPH VEDSFEKNNG GDLISSNFLP NYSDGYKGGQ 240 LPGPGLDYVP LVQQGAGTND YNQFRTLDLA SWEEVFQQCT RGSGDIPANP SISGAPLLGS 300 TQNFSESFSQ LLDAGFGVNS LSGGSISPYL QSNNPLEPSS LEYQDNANVS VTAKKPVLGN 360 IRGEEGLQKV DSFSKWMTKE LGGVDDLHLK SSSNMSWQNI ESATAVDDHS IELENYSMSP 420 SIGQDQLFDI KDFSPSCAST DGETKVVITG KFLVSPSEVF KYSWACMFGE VEVPAEVLGN 480 GVLCCYAPLH SAGRVPFYVT CSNRFACSQV REFEYLVESQ NANSANVCGN GMTEKQLCLR 540 LQKLLALKSP GNFESTSENI RKKEQIFRKI LLLMEDELCL GVEQHLKEKV VSWLFDKVSD 600 DGKGPNILDE EGQGFIHLAA ALGFDWIIPP TIAAGVSVNF RDINGWTALH WAAFYGREKT 660 VALLVALEAA SGALTDPSPE FPLGRTAADL ASANGYKGIS GFLAEHSLTA HLETLTMTDR 720 KSDSSLEDSM SKAVQTMREK VATPDDEGDG SDLSLKDSLA AVRNATQAAG RIHQVFRMQS 780 FQRKQVTKGA GDDESLLSDE QLLSLVSSRI QRPGRPDEQS HSAAVHIQKK FRGWKKRKEF 840 LIIRERVVKI QAHVRGHQVR KRYKTVVWSV GILEKVILRW RRKGTGLRGF RPDALNKASS 900 PHVAPVVQKK PSEEDDYDYL KEGRKQTEER LQKALVRVKS MVQYPEGRAQ YRRLLTAVEG 960 FQKNQQHVND NVISSSENVP VEGEDDMIDV ESLFDDDNFM AIAFE |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 908 | 924 | KKPSEEDDYDYLKEGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
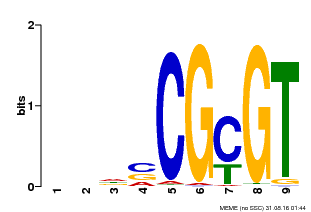
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010690491.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X1 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | A0A0K9QYQ2 | 0.0 | A0A0K9QYQ2_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010690491.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 0.0 | ethylene induced calmodulin binding protein | ||||



