 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv6_142620_qrgc.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 399aa MW: 44192.9 Da PI: 9.6694 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.2 | 9.9e-14 | 75 | 123 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+a++ a+++++g
Bv6_142620_qrgc.t1 75 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDVAAQRFRG 123
89****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 91.7 | 5.6e-29 | 213 | 329 | 1 | 94 |
EEEE-..-HHHHTT-EE--HHH.HTT.......................---.....--SEEEEEETTS-EEEEEE..EEETTEEEE-TTH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......................ggkke...esktltledesgrsWevkliyrkksgryvltkGW 65
f+kv tpsdv+k++rlv+pk++ae+h +++ l +ed + ++W+++++y+++s++yvltkGW
Bv6_142620_qrgc.t1 213 FEKVVTPSDVGKLNRLVIPKQHAEKHfplqgsnsttttttttsnttntlV---GsgsKGVLLNFEDVNQKVWRFRYSYWNSSQSYVLTKGW 300
99***************************977666666665555555441...0444788889999999********************** PP
HHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 66 keFvkangLkegDfvvFkldgrsefelvv 94
++Fvk++ Lk+gD+v F+++ +++l++
Bv6_142620_qrgc.t1 301 SRFVKEKSLKAGDIVSFQRSTGPDRQLYI 329
******************87656666655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 8.62E-27 | 75 | 130 | No hit | No description |
| Pfam | PF00847 | 3.2E-9 | 75 | 123 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.77E-17 | 75 | 131 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 20.455 | 76 | 131 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-30 | 76 | 137 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 8.4E-21 | 76 | 131 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 5.9E-38 | 210 | 335 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.05E-28 | 212 | 323 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.38E-25 | 212 | 320 | No hit | No description |
| Pfam | PF02362 | 3.2E-26 | 213 | 328 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.506 | 213 | 335 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 7.9E-23 | 213 | 333 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 399 aa Download sequence Send to blast |
MEASSMDESS STSDSQVSTT LPPMIVRNMM MVPSAPLPTK ASATAGSLCR MGSGTSVIID 60 SENGIEAESR KLPSSKYKGV VPQPNGRWGA QIYEKHQRVW LGTFNEEDEA ARAYDVAAQR 120 FRGRDAVTNF KPLSSEDELG AAELRFLNSR SKAEVVDMLR KHTYLDELQQ SRRNFKGDVC 180 NNFGEKGLLH SSSDKSKLSG NNNEGGFGRE QLFEKVVTPS DVGKLNRLVI PKQHAEKHFP 240 LQGSNSTTTT TTTTSNTTNT LVGSGSKGVL LNFEDVNQKV WRFRYSYWNS SQSYVLTKGW 300 SRFVKEKSLK AGDIVSFQRS TGPDRQLYID YKQRSISVEC TPVDVRAAGQ MIRLFGVDIC 360 KLPISVAEVT VNVNGKRLRE MELLAMESSS KKQMVVAAL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 7e-55 | 210 | 335 | 11 | 118 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
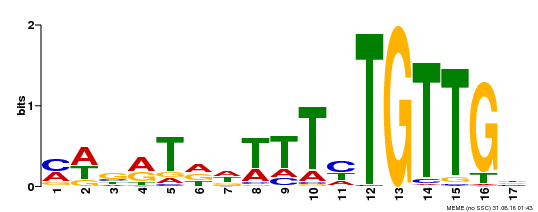
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010681667.1 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9ZWM9 | 1e-140 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | A0A0J8C877 | 0.0 | A0A0J8C877_BETVU; Uncharacterized protein | ||||
| STRING | XP_010681667.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-135 | related to ABI3/VP1 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



