 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv5_118270_wwos.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 996aa MW: 110703 Da PI: 5.808 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.6 | 1.4e-41 | 141 | 217 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
Cqve+C adl++ak+yhrrhkvCe+hska+++lv++++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+q+
Bv5_118270_wwos.t1 141 CQVEDCGADLTKAKDYHRRHKVCEMHSKATKALVANVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNKRRRKTQP 217
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 9.9E-34 | 133 | 202 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.111 | 138 | 215 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.44E-38 | 139 | 220 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.4E-30 | 141 | 214 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-9 | 733 | 887 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.09E-9 | 767 | 886 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.45E-9 | 780 | 887 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 996 aa Download sequence Send to blast |
MAEFHGIDPS QFRGMVKRGV EWDLNQWNWD GDLFIATPTN FQAKQFIPVV PVSGNSSNTS 60 SSCSDDVHYP IDDGRRELER KRRVIVVEED NLEEPGSLTL KLGGNCFPPV PEGEREVGNW 120 EGSSGKKTKL AAGGSSNRAL CQVEDCGADL TKAKDYHRRH KVCEMHSKAT KALVANVMQR 180 FCQQCSRFHA LQEFDEGKRS CRRRLAGHNK RRRKTQPDTV AGGNSVNDEQ TNSALLMSLL 240 RILSNMHANN KSNQSTDQDL VAQLLRSLAS PSGSHMGKGL HGVSHESQKI INGGIANKNG 300 HSEKMSHFLS NDDMNPPQVI DQHPAPPDSE IPRKGLYADN SKSSEIQAVS SQEPGSSFPM 360 KDSPPAYSEI TAGRMKLNNF DLNDAYVDSD DGMEDLERSP VDGNFVTGSA DYPSWAQQGS 420 HQSSPPQTSG NSDSASAQSP SSSSGEAQSR TDRIVFKLFG KEPGGSPIML RGEILEWLAH 480 SPTDIESYIK PGCIILTVYL RLAESLWEEL CSDLSSCLTR LFDISDDTFW RTGWVYVRVQ 540 NQIAVVHNGH VLLDTSLSLH NSNCSRILSV MPIAVSMDER ARFEVRGFNL SESTTRLLCA 600 MEGKYIDQEA TQESKDDCDF VEEDDEAEHA SIVCSVPNVT GRGFIEVEDP GLSTSFFPFI 660 VAEKDVCSEI RTLESVLGLK KSDEDAYEVN KTLESRRQAL DFINEMGWLL HRSHLKLRLA 720 DLDPNATIFS FRRFTWLMEF SMEHDWCAVV KKLLDVMVAG TVGSGEHPSL KVALLEMGLL 780 HRAVRRNSRS MVELLLEYVP MKFCDEFMSP VDGGKERFLF RPDAQGPAGL TPLHIAAGRD 840 ASEDVLDALL DDPGKIGLDA WKNAQDSTGA TPEDYARMRG HYAYIHMVQR KIQRSLASEH 900 VLDIPVEPSV APRHDGAVSF EVGTSVNQSC KLCDQKKMFM YYRSRVRTSL AYRPTMLSMV 960 GIAAVCVCVA LLFKSMPNVV CLFQPFRWEL LNYGSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-32 | 131 | 214 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
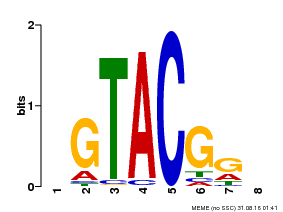
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00060 | PBM | Transfer from AT3G60030 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010678887.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A0J8BDT7 | 0.0 | A0A0J8BDT7_BETVU; Uncharacterized protein | ||||
| STRING | XP_010678887.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



