 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv4_095720_cndg.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 999aa MW: 110767 Da PI: 5.7114 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.4 | 1.5e-41 | 145 | 222 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adl++ak+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+q+
Bv4_095720_cndg.t1 145 VCQVEDCGADLTKAKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTQP 222
6**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.3E-33 | 139 | 207 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.206 | 143 | 220 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.96E-38 | 144 | 225 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.1E-29 | 146 | 219 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-8 | 750 | 890 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 7.77E-9 | 776 | 890 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 68 | 778 | 808 | IPR002110 | Ankyrin repeat |
| CDD | cd00204 | 5.47E-10 | 783 | 890 | No hit | No description |
| SMART | SM00248 | 0.94 | 831 | 861 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 999 aa Download sequence Send to blast |
MEQARRFHAL GPSEMRGMGK RGSEWDLNDW KWDGDLFIAT RSGGALPSNY QGQQFIPVSG 60 NSSNSSTSCS DDVHSSIDDG RRESERKRRV IVVEEDSLEE AGSLTLKLGG NCFPVSESER 120 EVGTWEGTSG KKTAKLVGGS SNRAVCQVED CGADLTKAKD YHRRHKVCEM HSKASKALVG 180 NVMQRFCQQC SRFHVLQEFD EGKRSCRRRL AGHNKRRRKT QPDTAVHGNS INDEQTNNVL 240 LMSLPRILSD MHANNKSDQS TDQDLVAQLL RSLANPSGLH VAKGLHGLLH ESHKLLNGGM 300 ANGNGHSEKM SDCLSNDHLN SPQVIDQHDA LPDSAIPRKE LYEANSRSSE MQVVSSQDPG 360 NLFPIKDSPP AHSETTAGRM KLNNFDLNDA YVDSDDGIED LERSPVDGHC TTGSVDFPSW 420 AGHVSHQSSP PQTNSDSASA QSPSSSSGEP QCRTDRIVFK LFGKEPGDFP MVLRGQILDW 480 LAHSPTDIES YIRPGCIILT IYLRLAESSW EELCSDLSSC LTRLFDISED TFWSSGWVYV 540 RVQNQIACFH NGQVILDTTL SLQNSNSGQI LSIMPIAVSM DERAEFEVRG FNLSGSTTRL 600 LCALEGKYLD QEATQESKED GESVEEDDET EHATMSCVIP KITGRGYIEV EDHGLSSSFF 660 PFIVAEKNVC SEIRTLESVL ELKEADEDAY EVKGTFESRR LALDFINEMG WLLHRSHLKS 720 RLADLDPNTV IFSFRRFTWL MDFSIDHDWC AVVRKLLDVL VDGTVGSGEH PSLKVAMSQM 780 GLLHKAVRKN SRSMVELLLK YVPDNLAEDF ISTADGGQVR FMFRPDAQGP AGLTPLHVAA 840 GRDGSEDVLD ALLDDPGKIG IDAWKNARDS TGATPEDYAR MRGQYAYIHM VQRKIQRSSA 900 ARHVVLDIPV ESSVSLRHDG AVSFEVGRNV TQSCKLCDQK MSVYYRSSAR TSLVYRPAML 960 SMVAIAAVCM CVALLFKSMP NVVCLFQPFR WESLNYGPS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 9e-32 | 139 | 219 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
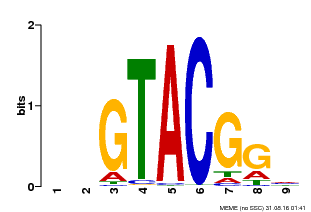
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010694791.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Refseq | XP_010694792.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A0J8BDT7 | 0.0 | A0A0J8BDT7_BETVU; Uncharacterized protein | ||||
| STRING | XP_010694791.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



