 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv4_074840_wwrw.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 623aa MW: 70135.7 Da PI: 6.5859 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.4 | 1.6e-13 | 570 | 618 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHT...TTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmg...kgRtlkqcksrwqky 47
r++WT+e ++ l ++v+++G++ Wk+I t Rt+ ++k++w++
Bv4_074840_wwrw.t1 570 RKKWTPEQEDALRNGVQKFGKR-WKLILDTYSslfRDRTDVDLKDKWRNM 618
789*******************.*************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.057 | 567 | 623 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.9E-9 | 569 | 621 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.4E-10 | 570 | 618 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.2E-13 | 570 | 620 | IPR009057 | Homeodomain-like |
| CDD | cd11660 | 1.77E-15 | 571 | 620 | No hit | No description |
| SuperFamily | SSF46689 | 1.48E-12 | 572 | 621 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 623 aa Download sequence Send to blast |
MDEKIARWIL DFLLRTSIDE KIINGVLSCL PLPNNDSRLK KTILLRKIES ETINGVVSEK 60 ILELLEIIEE LDHRMGKLAS DLMKEAYCAV AVDCTVRFLE ENVAENGKYF EAVERVWRNR 120 VCRMEGLISE DSNRWLRDIE AAVWDSGVCE KIWRRNTRNE ALKAVKGYVK DAWKCIGPTY 180 LELVANLVRD EGEELRRGSS DAGGETVGSD NPCLELVASE GMQRSLTTPR RKHVAIKRRR 240 LRGAKISDAD EVAEESNDIS NHKLDEAQEP LQLNSMELDA GVSETHPDVS RAAQNLSFSK 300 ARDSTSNVNE VENQTGPEIH AVPEVSRVQE APHSSNMEPH AVTDPLLETL STKATDTTSN 360 VKAVGNQTGL GMHATPEVIR VQEALNSSSR ELHAAVTDPL PKALQIATLA SRGVTVNKGT 420 GKENQTGSDT RAIPEAGNSS HDVFAAFEYV SSKMTTRNKR NGALLDNQKQ LGICAPREST 480 DNKSKNIEKK DDCTRDVLGN ETHDPPEQNK RKRSLMEPNG TSHTLEWGES ADSLNQDLDA 540 GPRSSLSQRR CRVSPLKQYE DKKFPWRRER KKWTPEQEDA LRNGVQKFGK RWKLILDTYS 600 SLFRDRTDVD LKDKWRNMMR YHS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 229 | 239 | RRKHVAIKRRR |
| 2 | 236 | 240 | KRRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
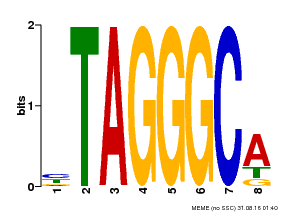
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010673889.1 | 0.0 | PREDICTED: uncharacterized protein LOC104890194 | ||||
| TrEMBL | A0A0K9QSX4 | 0.0 | A0A0K9QSX4_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010673889.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G15720.1 | 1e-18 | TRF-like 5 | ||||



