 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bostr.25542s0071.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Boechereae; Boechera
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 339aa MW: 37885.1 Da PI: 9.3451 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 312.7 | 1.2e-95 | 44 | 338 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yye................paaslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaer 70
m++ g++e r k+ yy+ ++++l +++++ms++aerda+++ern a+++kk+a+a+rd+a+ qrdkal+er
Bostr.25542s0071.1.p 44 MESGGQYEngRCKSdYYKgtqsvwnmmpqhqmkeQHNALV--MNKKIMSILAERDAAVKERNEAVAAKKEALAARDEALEQRDKALSER 130
7888888878999889999999999997766644445555..99********************************************* PP
GAGA_bind 71 nkalverdnkllalllvenslasalpvgvqvlsgtksidslqqlsepqledsavelreeeklealpieeaaeeakekkkkkkrqrakkp 159
++a++er+++l al+++en+l+ l s +k ++ ++e++ p+++ ++ea +++ +k r+k+
Bostr.25542s0071.1.p 131 DNAIMERESALNALQYRENNLNYIL-------SCAKRGG-----------SQSCVTEESHLPAPSPVSTIPPEAANTRPTK---RKKES 198
*******************965443.......2222221...........1122234566667889999999999999888...45555 PP
GAGA_bind 160 kekkakkkkkksekskkkvkkesaderskaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvst 248
k+ ++k kk +e+ + +v+++++ +s+++++s d++ln v++De+t+PvP C+CtG+ rqCYkWG GGWqS+CCttt+S+yPLP+++
Bostr.25542s0071.1.p 199 KQGARSKGKKVGEDLNCQVASPGM--KSRKDWDSNDVGLNLVTFDETTMPVPRCTCTGTARQCYKWGHGGWQSSCCTTTLSQYPLPQMP 285
666666777778899999999995..6************************************************************** PP
GAGA_bind 249 krrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
++r++Ri+grKmS+++f++lL++La+eG+dls pvDLkd+WAkHGtn+++ti+
Bostr.25542s0071.1.p 286 NKRHSRIGGRKMSGSVFSRLLSRLAGEGHDLSSPVDLKDYWAKHGTNRYITIK 338
****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 4.9E-162 | 44 | 338 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.2E-92 | 61 | 338 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MLALDPHLPP CSLLRYGHHL HLILRDLSLS NSVDCFSYEI LKRMESGGQY ENGRCKSDYY 60 KGTQSVWNMM PQHQMKEQHN ALVMNKKIMS ILAERDAAVK ERNEAVAAKK EALAARDEAL 120 EQRDKALSER DNAIMERESA LNALQYRENN LNYILSCAKR GGSQSCVTEE SHLPAPSPVS 180 TIPPEAANTR PTKRKKESKQ GARSKGKKVG EDLNCQVASP GMKSRKDWDS NDVGLNLVTF 240 DETTMPVPRC TCTGTARQCY KWGHGGWQSS CCTTTLSQYP LPQMPNKRHS RIGGRKMSGS 300 VFSRLLSRLA GEGHDLSSPV DLKDYWAKHG TNRYITIK* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00480 | DAP | Transfer from AT4G38910 | Download |
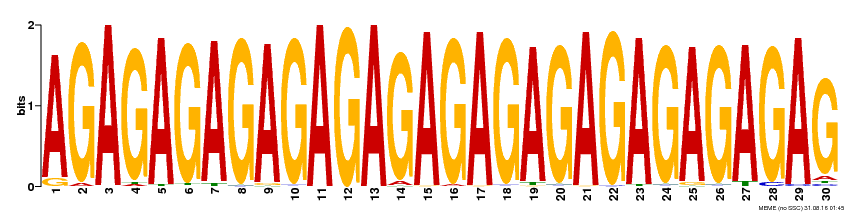
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bostr.25542s0071.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF633603 | 0.0 | EF633603.1 Cardamine pratensis GAGA-motif binding transcriptional activator (BBR/BPC5) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006411681.1 | 0.0 | protein BASIC PENTACYSTEINE5 | ||||
| Refseq | XP_024005542.1 | 0.0 | protein BASIC PENTACYSTEINE5 | ||||
| Refseq | XP_024005543.1 | 0.0 | protein BASIC PENTACYSTEINE5 | ||||
| Refseq | XP_024005544.1 | 0.0 | protein BASIC PENTACYSTEINE5 | ||||
| Swissprot | F4JUI3 | 0.0 | BPC5_ARATH; Protein BASIC PENTACYSTEINE5 | ||||
| TrEMBL | V4M9X7 | 0.0 | V4M9X7_EUTSA; Uncharacterized protein | ||||
| STRING | Bostr.25542s0071.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4090 | 26 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G38910.2 | 1e-175 | basic pentacysteine 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bostr.25542s0071.1.p |



