 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bostr.13671s0120.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Boechereae; Boechera
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 346aa MW: 38607.3 Da PI: 9.8904 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 42.4 | 1.7e-13 | 61 | 109 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+a++ a ++++g
Bostr.13671s0120.1.p 61 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARAYDVAVNRFRG 109
89****9888.8*********3.....5**********99************998 PP
| |||||||
| 2 | B3 | 101.2 | 6e-32 | 189 | 292 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f+k tpsdv+k++rlv+pk++ae+h ++++ +++ l +ed +g++W+++++y+++s++yvltkGW++Fvk+++L++gD+v F+
Bostr.13671s0120.1.p 189 FEKPVTPSDVGKLNRLVIPKHHAEKHfplpsSSVSVKGVLLNFEDVNGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKNLRAGDVVSFSK 277
89****************************9888889***************************************************6 PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
++ ++++l++++ ++
Bostr.13671s0120.1.p 278 SNGQDQQLYIGWKSR 292
655999999998776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.2E-8 | 61 | 109 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.17E-17 | 61 | 117 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 3.00E-26 | 61 | 119 | No hit | No description |
| PROSITE profile | PS51032 | 20.099 | 62 | 117 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.3E-28 | 62 | 123 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.6E-20 | 62 | 118 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 5.9E-42 | 181 | 294 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.27E-33 | 184 | 293 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.26E-29 | 187 | 280 | No hit | No description |
| PROSITE profile | PS50863 | 14.254 | 189 | 293 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.4E-25 | 189 | 293 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 7.1E-29 | 189 | 292 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 346 aa Download sequence Send to blast |
MEVSSVDEST TSTGSICETP AITPAKKSSV GNNLYRMGSG SSVVLDSENG IEAESRKLPS 60 SKYKGVVPQP NGRWGAQIYE KHQRVWLGTF NEEDEAARAY DVAVNRFRGR DAVTNFKDVR 120 MDEDEGEFLI SHSKSEIVDM LRKHTYNEEL EQSKRRRNGN GNTVRTMLTS GLSNDGVSTT 180 EFRSAEPLFE KPVTPSDVGK LNRLVIPKHH AEKHFPLPSS SVSVKGVLLN FEDVNGKVWR 240 FRYSYWNSSQ SYVLTKGWSR FVKEKNLRAG DVVSFSKSNG QDQQLYIGWK SRSGSDLVAG 300 RVLRLFGVNI SPGSLRNDVV GNKRVNDTEM ISLVCSKKQR IFHAS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 1e-74 | 185 | 299 | 10 | 124 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
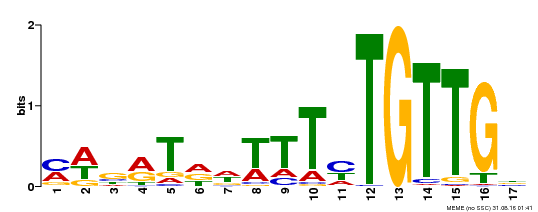
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bostr.13671s0120.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB013886 | 0.0 | AB013886.1 Arabidopsis thaliana mRNA for RAV1, complete cds. | |||
| GenBank | AC011810 | 0.0 | AC011810.8 Arabidopsis thaliana chromosome I BAC T6J4 genomic sequence, complete sequence. | |||
| GenBank | AY063855 | 0.0 | AY063855.1 Arabidopsis thaliana putative DNA-binding protein RAV1 (At1g13260) mRNA, complete cds. | |||
| GenBank | AY091291 | 0.0 | AY091291.1 Arabidopsis thaliana putative DNA-binding protein RAV1 (At1g13260) mRNA, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010476321.1 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9ZWM9 | 0.0 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | D7KQD7 | 0.0 | D7KQD7_ARALL; Uncharacterized protein | ||||
| STRING | Bostr.13671s0120.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 0.0 | related to ABI3/VP1 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bostr.13671s0120.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



