 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bostr.0124s0004.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Boechereae; Boechera
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 837aa MW: 91705.2 Da PI: 6.2763 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Bostr.0124s0004.1.p 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 167.1 | 1.2e-52 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..E CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..g 89
+aee+++e+++ka+ ++ W +++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+++d++ W + ++++++++v+ ++ g
Bostr.0124s0004.1.p 161 IAEETLAEFISKATGTAVEWIQMPGMKPGPDSIGIIAISHGCTGVAARACGLVGLEPT-RVAEIVKDRPSWFRECRAVDVMNVLPTAngG 249
7899******************************************************.7777777777****************9999* PP
EEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SS CS
START 90 alqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgr 175
+++l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ s+q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl++
Bostr.0124s0004.1.p 250 TIELLYMQLYAPTTLAPpRDFWLLRYTSVLEDGSLVVCERSLKSTQNGPSmplVQNFVRAEMLPSGYLIRPCDGGGSIIHIVDHMDLEAC 339
*************************************************999999*********************************** PP
XXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 176 lphwllrslvksglaegaktwvatlqrqce 205
+++++lr+l++s + ++kt++a+l+++++
Bostr.0124s0004.1.p 340 SVPEVLRPLYESPKVLAQKTTMAALRQLKQ 369
**************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.71E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 9.27E-17 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 2.9E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.273 | 20 | 76 | IPR001356 | Homeobox domain |
| CDD | cd14686 | 1.29E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 25.363 | 151 | 379 | IPR002913 | START domain |
| CDD | cd08875 | 1.38E-73 | 155 | 371 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.0E-20 | 160 | 346 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 1.65E-35 | 160 | 372 | No hit | No description |
| SMART | SM00234 | 2.4E-36 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 3.2E-50 | 161 | 369 | IPR002913 | START domain |
| Pfam | PF08670 | 8.7E-52 | 693 | 835 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 837 aa Download sequence Send to blast |
MAMSCKDGKL GCLDNGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VHENSYFRQN 120 THNPSLPAKD TSCESVVTSG QHELASQNPQ RDASPAGLLS IAEETLAEFI SKATGTAVEW 180 IQMPGMKPGP DSIGIIAISH GCTGVAARAC GLVGLEPTRV AEIVKDRPSW FRECRAVDVM 240 NVLPTANGGT IELLYMQLYA PTTLAPPRDF WLLRYTSVLE DGSLVVCERS LKSTQNGPSM 300 PLVQNFVRAE MLPSGYLIRP CDGGGSIIHI VDHMDLEACS VPEVLRPLYE SPKVLAQKTT 360 MAALRQLKQI AQEVTQTNSS VNGWGRRPAA LRALSQRLSR GFNEAVNGFT DEGWSVIGDS 420 MDDVTITVNS SPDKLMGLNL TFSNGFAPVS SVVLCAKASM LLQNVPPAIL LRFLREHRSE 480 WADNNIDAYL AAAVKVGPCS ARVGGFGGQV ILPLAHTIEH EEFMEVIKLE GLGHSPEDAI 540 VPRDIFLLQL CSGMDESAVG TCAELIFAPI DASFADDAPL LPSGFRIIPL DSAKEVSSPN 600 RTLDLASALE IGSAGTKAST DQSGNSTCAR SVMTIAFEFG IESHMQEHVA SMARQYVRGI 660 ISSVQRVALA LSPSHISSQV GLRTPLGTPE AQTLARWICQ SYRGYMGVEL LKSISEGNES 720 ILKNLWHHTD AIICCSMKAL PVFTFANQAG LDMLETTLVA LQDISLEKIF DDNGRKTLCS 780 EFPQIMQQGF ACLQGGICLS SMGRPVSYER AVAWKVLNEE ENAHCICFVF ISWSFV* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
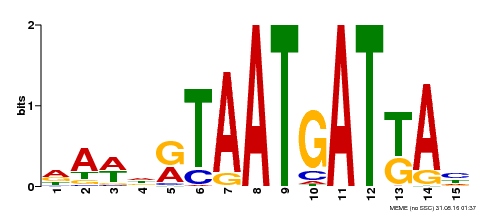
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bostr.0124s0004.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ439449 | 0.0 | AJ439449.1 Arabidopsis thaliana mRNA for homeodomain-leucine zipper protein (ATHB-15 gene). | |||
| GenBank | AY060556 | 0.0 | AY060556.1 Arabidopsis thaliana At1g52150/F5F19_21 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020868586.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | D7KIP0 | 0.0 | D7KIP0_ARALL; ATHB-15 | ||||
| STRING | Bostr.0124s0004.1.p | 0.0 | (Boechera stricta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2022 | 26 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bostr.0124s0004.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



