 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast07G188100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 474aa MW: 51612.2 Da PI: 5.5677 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 98.9 | 3.4e-31 | 266 | 320 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k rlrWt eLHe+Fveav++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYR
Brast07G188100.1.p 266 KSRLRWTLELHESFVEAVNKLDGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRH 320
68****************************************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 10.912 | 263 | 323 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.4E-16 | 264 | 320 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-29 | 264 | 321 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.9E-24 | 266 | 320 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.3E-7 | 268 | 319 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.2E-22 | 358 | 403 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 474 aa Download sequence Send to blast |
MSSQGVVAVK PTAAPDKTVH YACGSTQPSV HKLLDAKLDH LGLLDDNLSS TSQSSNIKTE 60 LIRSSSLPFR LQERSTQSDT ESPLSQVSHP NFSDPMVSNS STFCTSLFSS SSTNSSPCQQ 120 MGALPFLPHP PKCEQQVILG QSCSSSLHLS GGDIGNDYDE AEQSDDMKDF LNLPVDASDA 180 SYHGENNAMA FTEQMEFQFL SEQLGIAITD NEESPLLDDI YDTPPPQLSS LPVSSCSNQS 240 LHNLGSPVKL PLSSARSSSG STTANKSRLR WTLELHESFV EAVNKLDGPE KATPKGVLKL 300 MKVEGLTIYH VKSHLQKYRH ARYLPDTKED KKATLDCKKV QSAQSGSNGS DLDKNKNLAE 360 ALRMQMEVQK QLHEQLEVQR QLQLRIEEHA KYLQRILEEQ QKTSNGGSSS LKISTEQPES 420 TSKNGTSPEE ATTSSPHPSK NIAPSDAGTE CDSPVRNKRA RVHGDLESES PCS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 3e-27 | 266 | 323 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling. Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes. Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis. {ECO:0000250|UniProtKB:Q6YXZ4}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
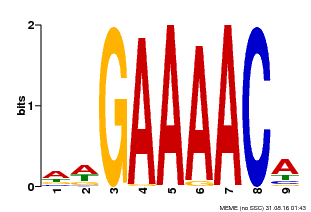
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast07G188100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003563523.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_024313685.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Swissprot | A2X0Q0 | 1e-155 | PHR3_ORYSI; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | I1GWS3 | 0.0 | I1GWS3_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G34470.4 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 4e-64 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast07G188100.1.p |



