 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast04G330100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 867aa MW: 96796.9 Da PI: 7.0743 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.6 | 1.1e-05 | 559 | 580 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF + +Lk+H+++H
Brast04G330100.1.p 559 TCEECGASFQKPAHLKQHMQSH 580
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 20.3 | 1.5e-06 | 586 | 610 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s+ rk++L+rH+ +H
Brast04G330100.1.p 586 FACPleDCPFSYIRKDHLNRHMLKH 610
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 14.3 | 0.00012 | 615 | 640 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ Cp C+k+Fs k n +rH + H
Brast04G330100.1.p 615 FACPmeGCDKRFSIKANVQRHVKEmH 640
78********************9988 PP
| |||||||
| 4 | zf-C2H2 | 13.8 | 0.00017 | 654 | 676 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
C+ C+k F+ s Lk+H +H
Brast04G330100.1.p 654 CKeeGCNKAFKYASKLKKHEESH 676
88889**************9988 PP
| |||||||
| 5 | zf-C2H2 | 18.3 | 6.2e-06 | 743 | 767 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC sFs+ksnL++H++ H
Brast04G330100.1.p 743 KCTfeDCESSFSNKSNLTKHMKAcH 767
79999*****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.25.40.10 | 1.6E-7 | 32 | 113 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.864 | 35 | 65 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.761 | 66 | 101 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.19 | 68 | 97 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.309 | 137 | 167 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 4.5E-4 | 142 | 164 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 9.8E-4 | 142 | 164 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.583 | 168 | 198 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.01 | 170 | 198 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 10.315 | 199 | 233 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 2.6E-5 | 201 | 230 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 2.2E-5 | 201 | 234 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.859 | 269 | 299 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.6E-7 | 272 | 331 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF01535 | 0.0059 | 274 | 295 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF13041 | 2.7E-11 | 299 | 347 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 12.244 | 300 | 334 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 8.5E-9 | 302 | 336 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.32 | 335 | 365 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.369 | 371 | 401 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.06 | 374 | 398 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.6E-7 | 401 | 465 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.426 | 403 | 433 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.136 | 437 | 471 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 33 | 512 | 534 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.757 | 558 | 585 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0088 | 558 | 580 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.2E-4 | 559 | 576 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 560 | 580 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.24E-10 | 570 | 612 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.4E-11 | 577 | 612 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.016 | 586 | 615 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.2E-4 | 586 | 610 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 588 | 610 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.21E-7 | 596 | 644 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 8.3E-6 | 613 | 639 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0012 | 615 | 640 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.637 | 615 | 645 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 617 | 640 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-6 | 650 | 676 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.026 | 652 | 676 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.471 | 652 | 676 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 654 | 676 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7.4 | 684 | 709 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 686 | 709 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 712 | 733 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.1E-6 | 723 | 764 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.0E-8 | 724 | 784 | No hit | No description |
| PROSITE profile | PS50157 | 11.884 | 742 | 772 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 742 | 767 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 744 | 767 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.7E-5 | 767 | 795 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.4 | 773 | 799 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.891 | 773 | 799 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 775 | 799 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 867 aa Download sequence Send to blast |
MNRSICHHLL AHGKSIRELE QIHAAALVHG LHPGNQSVSC KMFRSYADLG RAADARRLFG 60 EIPKPDLVSL TSLMSLHLQL NQHREAMSLF SRVVASGGHR PDGFAVVGAL SASGGVGDLR 120 AGEALHGMIC RCGLGSEPVV GNALVDMYSR CGRFEGAVGV FDEMALKDEI TWGSMLHGYI 180 KCVGVDSALS FFDQMPMKSV VSWTALITGH VQAKNPIRAL ELFGRMVLEG HRPTHFTIVG 240 VLSACADIGA LDLGRVIHGY GSKFNVSTNI IVSNALMDMY AKSGSIEMAF AVFEEVPLKD 300 AFTWTTMISS FTVQGDGSKA LELFGDMLRS GIVPNSVTFV SVLSACSHAG LIQNGKELFD 360 KMRGTYNISP QLEHYGCMVD LLGRGGLLEE AEALIYDMDL EPDVVIWRSL LSACLVHGND 420 RLAEIAGKEI IKREPGDDGV YVLLWNIYAS SDKWKEALDM RKQMLTRRVY KKPGCSWIEV 480 DGGVHEFLMS SGDEIDGDEM VEETHGRDIR RYKCEFCTVI RSKKCLIRAH MVAHHKDELD 540 MSEIYNSNGE KIIQEGEPTC EECGASFQKP AHLKQHMQSH SKERSFACPL EDCPFSYIRK 600 DHLNRHMLKH EGKLFACPME GCDKRFSIKA NVQRHVKEMH EVENGCKSNQ QFLCKEEGCN 660 KAFKYASKLK KHEESHVKLD YVEVVCCEEG CMKTFTNVEC LRAHNQSCHQ YVQCDICGEK 720 HLKKNIKRHL RAHDTMVPTE RIKCTFEDCE SSFSNKSNLT KHMKACHDQI RPFACRFEGC 780 GKEFTYKHVR DNHEKSSAHV YVEGDFAEMA EQLRSCPRGG RKRAAVTVET LTRKRVSMPG 840 EAVSLEDGTE YMRWLLSGGD VSSHAE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 5e-58 | 20 | 414 | 4 | 331 | PLS9-PPR |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
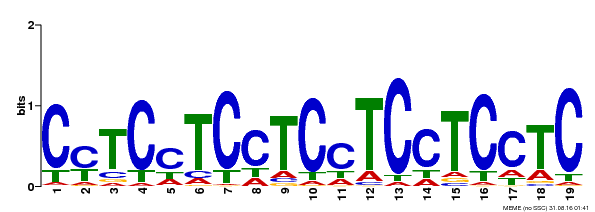
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast04G330100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024317701.1 | 0.0 | pentatricopeptide repeat-containing protein At2g22410, mitochondrial-like | ||||
| TrEMBL | A0A453MVM8 | 0.0 | A0A453MVM8_AEGTS; Uncharacterized protein | ||||
| STRING | BRADI3G01487.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-114 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast04G330100.1.p |



