 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast03G129400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 426aa MW: 46761.6 Da PI: 8.8344 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.5 | 1.2e-05 | 92 | 114 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++C+k F r nL+ H r H
Brast03G129400.1.p 92 FVCEICNKGFQRDQNLQLHRRGH 114
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.3 | 0.00025 | 179 | 201 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC+ Cgk++ +s+ k H +++
Brast03G129400.1.p 179 WKCERCGKRYAVHSDWKAHVKNC 201
58******************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.97E-8 | 91 | 114 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 91 | 114 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF12171 | 6.5E-5 | 92 | 114 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE profile | PS50157 | 10.949 | 92 | 114 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.017 | 92 | 114 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 94 | 114 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 96 | 144 | 174 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-4 | 167 | 199 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.97E-8 | 174 | 199 | No hit | No description |
| SMART | SM00355 | 93 | 179 | 199 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 426 aa Download sequence Send to blast |
MMLSDLSSDQ EATGSNSHGG GGDVVLGNQH VLLSPLFAPP SQALLLRPAP QLEQLQQPMA 60 GGKRKRSQPG NPDPGAEVIA LSPRALVATN RFVCEICNKG FQRDQNLQLH RRGHNLPWKL 120 RQRGSLPAAH NKQGGDGGPR KRVYVCPEPS CVHHDPARAL GDLTGIKKHF SRKHGEKRWK 180 CERCGKRYAV HSDWKAHVKN CGTREYRCDC GILFSRKDSL LTHRAFCDAL AEESARILAA 240 ANNSTNSRNS NIISNSNSGS RSNNNLLSPP PLFPTFSSHA QNPNPLTFLS QEPRYHHHHQ 300 QQEEELLPPF QPLYSYLDHV DLPMSAAITG SVSAFDADRD TLSFGLTPEG SVTMHAGGRR 360 RLTRDFLGAG EVMEDQLQMP LCSANAAAYQ GRHVATAASV CCATDLTRQY LGRLPPVSET 420 WSHNF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 2e-33 | 175 | 244 | 3 | 72 | Zinc finger protein JACKDAW |
| 5b3h_F | 2e-33 | 175 | 244 | 3 | 72 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that acts as a flowering master switch in both long and short days, independently of the circadian clock (PubMed:18790997, PubMed:18725639, PubMed:18774969, PubMed:19304997, PubMed:24280027). Promotes flowering upstream of HD1 by up-regulating FTL1, FTL4, FTL5, FTL6, EHD1, HD3A and RFT1 (PubMed:18790997, PubMed:18725639, PubMed:18774969). Seems to repress FTL11 expression (PubMed:18725639). May recognize the consensus motif 5'-TTTGTCGTAAT-3' in target gene promoters (PubMed:18725639). {ECO:0000269|PubMed:18725639, ECO:0000269|PubMed:18774969, ECO:0000269|PubMed:18790997, ECO:0000269|PubMed:24280027, ECO:0000303|PubMed:19304997}. | |||||
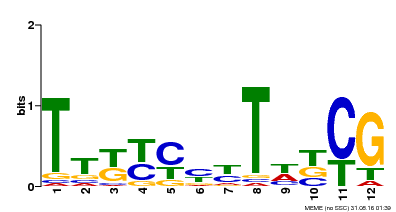
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00673 | SELEX | Transfer from GRMZM2G011357 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast03G129400.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by FKF1 (PubMed:25850808). Down-regulated by DLF1 (PubMed:25036785). During the basic vegetative growth phase (BVP, photoperiod-insensitive phase), suppressed via phytochrome-mediated light signals involving the phytochromobilin synthase HY2 (PubMed:25573482). {ECO:0000269|PubMed:25036785, ECO:0000269|PubMed:25573482, ECO:0000269|PubMed:25850808}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010234737.1 | 0.0 | protein indeterminate-domain 12 isoform X1 | ||||
| Swissprot | B1B534 | 1e-154 | EHD2_ORYSJ; Protein EARLY HEADING DATE 2 | ||||
| TrEMBL | A0A0Q3HTA8 | 0.0 | A0A0Q3HTA8_BRADI; Uncharacterized protein | ||||
| STRING | Traes_6DS_DE865EA6D.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP13096 | 28 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G55110.1 | 8e-99 | indeterminate(ID)-domain 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast03G129400.1.p |



