 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast02G277700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 390aa MW: 41952.3 Da PI: 7.4043 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 147.1 | 1.2e-45 | 121 | 277 | 1 | 136 |
TCP 1 aagkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec.eaesssssasnsssg.. 88
aa++kdrhski+T++g+RdRR+Rls+++a++fF+Lqd+LGfdk+skt++WLl+ +k ai e + ++s+c e+++ sss s s+ +
Brast02G277700.1.p 121 AAARKDRHSKICTAGGMRDRRMRLSLDVARKFFALQDMLGFDKASKTVQWLLNTSKSAIREVMAP---HSSACeEDDDGSSSISLSNMPap 208
589************************************************************99...55567655555555553333355 PP
TCP 89 .....................kaaksaakskksqksaasalnlakesrakarararertrekmriknkl 136
+aa++ ++s++++ a+++ ++kesr+kar+rarert+ek+r+++++
Brast02G277700.1.p 209 ekttgddrgegkkkpraarssRAASNPKPSSRKSGGANAHTIPDKESRSKARERARERTKEKNRMRWVT 277
666777778888888999875555555566666666666666777*********************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 3.3E-44 | 123 | 273 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 34.798 | 124 | 182 | IPR017887 | Transcription factor TCP subgroup |
| PROSITE profile | PS51370 | 11.607 | 253 | 270 | IPR017888 | CYC/TB1, R domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048831 | Biological Process | regulation of shoot system development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 390 aa Download sequence Send to blast |
MFPFCDSPMD LPLYQQLQLS PPSPKPDPEE DHHHQSSFFY YHSSPAFAGS DADAAFHHNS 60 CYLDPGAAAA TLPSADIDFS PPPELSLMDQ APAAPHPTAG NTERHGSDSG AGALESSRAA 120 AAARKDRHSK ICTAGGMRDR RMRLSLDVAR KFFALQDMLG FDKASKTVQW LLNTSKSAIR 180 EVMAPHSSAC EEDDDGSSSI SLSNMPAPEK TTGDDRGEGK KKPRAARSSR AASNPKPSSR 240 KSGGANAHTI PDKESRSKAR ERARERTKEK NRMRWVTLAS TINLESADEL TMTSPPNSNI 300 LKYRSSSSSM NTAVSAKLED RCSSSTATNG GRTVQEASIA SHAIMAGAFG DGGIYGSSGS 360 GSNYYYQHQQ LEEQQWELGG VVFANSRLY* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that functions as a negative regulator of lateral branching, presumably through its expression in axillary buds (PubMed:12581309, PubMed:20547591). Involved in the fine tuning of shoot branching. May function as an integrator of multiple signaling pathways to regulate the development of axillary buds. Works partially downstream of strigolactones to inhibit bud outgrowth (PubMed:20547591). Binds to MADS57 to suppress the negative regulation of D14 by MADS57 and balance the expression of D14 for tillering (PubMed:23463009). {ECO:0000269|PubMed:12581309, ECO:0000269|PubMed:20547591, ECO:0000269|PubMed:23463009}. | |||||
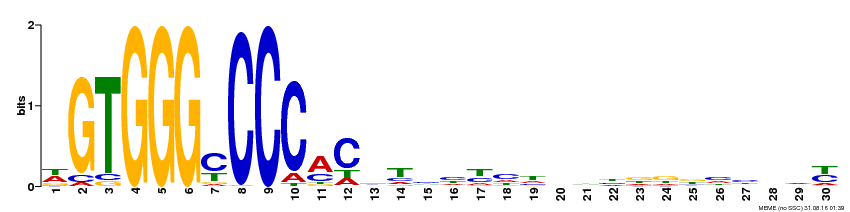
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00215 | DAP | Transfer from AT1G67260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast02G277700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ842222 | 5e-70 | DQ842222.1 Phyllostachys praecox TB1-1 protein (TB1-1) mRNA, complete cds. | |||
| GenBank | DQ842223 | 5e-70 | DQ842223.1 Phyllostachys praecox TB1-2 protein (TB1-2) mRNA, complete cds. | |||
| GenBank | DQ842225 | 5e-70 | DQ842225.1 Sasa fortunei TB1 protein (TB1) gene, partial cds. | |||
| GenBank | DQ910763 | 5e-70 | DQ910763.1 Yushania niitakayamensis TB1-like gene, complete cds. | |||
| GenBank | DQ910764 | 5e-70 | DQ910764.1 Pleioblastus amarus TB1-like gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003559513.1 | 1e-173 | transcription factor TEOSINTE BRANCHED 1 | ||||
| Swissprot | Q8LN68 | 5e-99 | TB1_ORYSJ; Transcription factor TB1 | ||||
| TrEMBL | I1GP36 | 1e-172 | I1GP36_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G11060.1 | 1e-172 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7253 | 36 | 51 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68800.1 | 4e-24 | TCP domain protein 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast02G277700.1.p |



