 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Brast02G074400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1034aa MW: 115605 Da PI: 5.9073 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 177.1 | 2.2e-55 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseen 91
lke ++rwl++ ei++iL+n+++++++ e+++rp sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een
Brast02G074400.1.p 19 LKEaQHRWLRPAEICEILKNYRNFRIAPEPPNRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKSGSIDVLHCYYAHGEEN 109
45559************************************************************************************** PP
CG-1 92 ptfqrrcywlLeeelekivlvhylevk 118
+fqrr+yw+Lee++ +ivlvhylevk
Brast02G074400.1.p 110 INFQRRSYWMLEEDFMHIVLVHYLEVK 136
************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.447 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-76 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.5E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.3E-5 | 444 | 531 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 0.00395 | 444 | 531 | No hit | No description |
| SuperFamily | SSF81296 | 4.76E-16 | 445 | 530 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 6.8E-6 | 445 | 529 | IPR002909 | IPT domain |
| CDD | cd00204 | 6.95E-13 | 628 | 737 | No hit | No description |
| SuperFamily | SSF48403 | 1.52E-16 | 636 | 740 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.4E-17 | 639 | 742 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.767 | 645 | 739 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.79 | 645 | 677 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.033 | 678 | 707 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.63 | 678 | 710 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1400 | 717 | 746 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 4.25E-8 | 849 | 902 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.12 | 851 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.663 | 852 | 881 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0034 | 853 | 872 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0013 | 874 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.487 | 875 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0017 | 876 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1034 aa Download sequence Send to blast |
MAEGRRYAIA PQLDIEQILK EAQHRWLRPA EICEILKNYR NFRIAPEPPN RPASGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKSGSIDVLH CYYAHGEENI NFQRRSYWML 120 EEDFMHIVLV HYLEVKAGKS SSRTREHDNM LQGARVDSPL SQLPSQTTDG ESSLSGQASE 180 YEETESDIYS GGAGYHSISR MQQHENGAGP IIDASFYSSY VPASSVGNQG LQATTTNTGF 240 YSYDQDNLPV VPNESGHGTP FNGPNGQFDL SSWNEMTKPD KRIHQMPPYG THVPSEHSPF 300 TEGPGIESFT FDEVYSNGLG IKDNSHADTD AEPLWQLPSA IGGSFATVDS FQQINGFLEE 360 AINYPLLKTQ SSNLSDILKD SFKKSDSFTR WMTKELADVD DSEIKPSSEY WNSEDADNII 420 GTSSHDQLDQ FTLGPMLAQD QLFSIIDFSP SWAYAGAKTR ILVTGKFLKP DEVIRLKWSC 480 MFGEIEVPAE ILADGTLGCY SPSQKTGRVP FYVTCSNRLA CSEVREFEYR PSNSQYMDAP 540 SLHGARNKTC LQMRLDKLLS LGPDEFHATL SNNTKELIDL NRKISLLMMN NDSWSELLKL 600 ADDNELITDD KQDQFLENCI RDKLHIWLLH KAGDGGKGPS VLDTEGQGVL HLAAALGYDW 660 AIRPTITAGV NINFRDARGW TALHWAAFCG RERTVVALIA LGAAPGALTD PSPDFPSGST 720 PADLASSNGH KGISGFLAEF SLTSHLQTLN LKEAMGSNAS EISGLPGIGD VTERSASPSA 780 PEGLQTGSMG DSLGAVRNAA QAAARIYQVF RVQSFQRKQA VQYEDDNGMI SDERALSLLS 840 YKTSKPGQFD PKHAAATRIQ NKFRGWKGRK EFLLLRRRVV QIQAHVRGHQ VRKHYRKIIW 900 SVGIVEKVIL RWRRRGAGLR GFRSTEGAPD STSSSAVDVI PNKPGEDDYS FLQEGRKQTE 960 ERLQRALARV KSMVQYPDAR DQYQRILTVV TKMQESQPMQ ENVLEESTEM DEGFLMSEFQ 1020 ELWDDDTPMP GYF* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
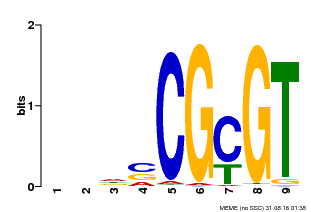
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Brast02G074400.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003558617.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| TrEMBL | I1H8R1 | 0.0 | I1H8R1_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G71810.2 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Brast02G074400.1.p |



