 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009149907.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 427aa MW: 46956.4 Da PI: 8.7794 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 125.1 | 2.2e-39 | 105 | 165 | 2 | 62 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkkss 62
++k+l+cprC+s++tkfCyynny+++qPr+fCkaC+ryWt+GG++rnvPvG+grrk+k+ss
XP_009149907.1 105 PTKILPCPRCKSMDTKFCYYNNYNINQPRHFCKACQRYWTAGGTMRNVPVGAGRRKHKSSS 165
67899*****************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 7.0E-31 | 81 | 162 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 6.1E-32 | 107 | 163 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.435 | 109 | 163 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 111 | 147 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 427 aa Download sequence Send to blast |
MMMESRDPAI KLFGMKIPFP AVFEPTTAVA LEEDYSGGDD TSPEKVTTEQ ATPEKKNNCN 60 NKSLNNSNDS KPETGDKEEA TSTDQIESDE TNQQTTADGK TLKKPTKILP CPRCKSMDTK 120 FCYYNNYNIN QPRHFCKACQ RYWTAGGTMR NVPVGAGRRK HKSSSSQYRH ITISEALQAA 180 RLDPGLQANT RVLSFGLQAP HQQHAAPMTP VMKLQGDQKV SNGARNARVE NGDDCSSGSS 240 VTTSVDETRA QSCRVVEPQV NNNNMNGYAC IPGVPWPYTW NPAMPPPGFY PPPGYPMPFY 300 PYWTIPMAPP NQSSSPMSQK GSSPNSPTLG KHYRDEDSST ERKQRNGCVI VPKTLRIDDP 360 NEAAKSSIWT TLGIKNEGST LGSKGGGMFK GFDQKTNKSN KDQTNNSHVL SANPAALSRS 420 LNFQERV |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.18278 | 0.0 | root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the vasculature of cotyledons and hypocotyls, leaves and roots. {ECO:0000269|PubMed:19619493}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
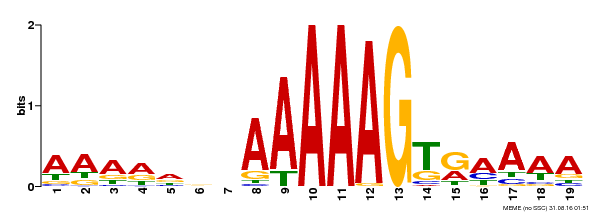
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Regulates a photoperiodic flowering response. Transcriptional repressor of 'CONSTANS' expression. {ECO:0000250, ECO:0000269|PubMed:19619493}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00394 | DAP | Transfer from AT3G47500 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009149907.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Highly expressed at the beginning of the light period, then decreases, reaching a minimum between 16 and 29 hours after dawn before rising again at the end of the day. {ECO:0000269|PubMed:19619493}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU451312 | 0.0 | GU451312.1 Brassica napus putative dof zinc finger protein (dof1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009149907.1 | 0.0 | PREDICTED: cyclic dof factor 3 | ||||
| Swissprot | Q8LFV3 | 0.0 | CDF3_ARATH; Cyclic dof factor 3 | ||||
| TrEMBL | A0A078JMT1 | 0.0 | A0A078JMT1_BRANA; BnaAnng22720D protein | ||||
| TrEMBL | M4DNQ4 | 0.0 | M4DNQ4_BRARP; Uncharacterized protein | ||||
| STRING | Bra018141.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5441 | 28 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47500.1 | 0.0 | cycling DOF factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



