 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009148531.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 341aa MW: 38088.5 Da PI: 9.4019 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 37.4 | 6.4e-12 | 58 | 106 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f ++eAa+ ++ a +++g
XP_009148531.1 58 SKYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEDEAARSYDVAVYRFRG 106
89****9888.8*********3.....5**********99***********9998 PP
| |||||||
| 2 | B3 | 101.1 | 6.5e-32 | 185 | 288 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
f+k+ tpsdv+k++rlv+pk++ae+h ++++ +++ l +ed +g++W+++++y+++s++yvltkGW++Fvk+++L +gD+v F++++ +++
XP_009148531.1 185 FEKAVTPSDVGKLNRLVIPKHHAEKHfplpsSNVSVKGVLLNFEDVTGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKNLLAGDVVSFSRSEGQDQ 279
89****************************9888889***************************************************9887999 PP
..EEEEE-S CS
B3 91 elvvkvfrk 99
+l++++ ++
XP_009148531.1 280 QLYIGWKSR 288
999999776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.39E-25 | 58 | 116 | No hit | No description |
| SuperFamily | SSF54171 | 2.42E-16 | 58 | 114 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 3.4E-7 | 58 | 106 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.257 | 59 | 114 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.2E-20 | 59 | 115 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.9E-28 | 59 | 120 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.3E-41 | 177 | 291 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 6.54E-32 | 183 | 291 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.20E-27 | 184 | 276 | No hit | No description |
| Pfam | PF02362 | 6.6E-29 | 185 | 288 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.2E-25 | 185 | 289 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 14.057 | 185 | 289 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MEVSSVDEST TSTGSICETP AMTSAKSSVN LHRMGSGSSV VLDSENGVEA ESRKLPSSKY 60 KGVVPQPNGR WGAQIYEKHQ RVWLGTFNEE DEAARSYDVA VYRFRGRDAV TNFKEARLDD 120 GEVEFLSSHS KAEIVDMLRK HTYSEELEQS KRRRNGNGNT ARTLTSSLNG DGVSTTEFRS 180 AEALFEKAVT PSDVGKLNRL VIPKHHAEKH FPLPSSNVSV KGVLLNFEDV TGKVWRFRYS 240 YWNSSQSYVL TKGWSRFVKE KNLLAGDVVS FSRSEGQDQQ LYIGWKSRSG PDVETGRVLR 300 LFGVNISPES SRTEVVGSKR VNDTEMLSLV CSKKQRIFHA S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 4e-74 | 181 | 295 | 10 | 124 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.17227 | 0.0 | leaf| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in all tissues examined: Roots, rosette leaves, cauline leaves, inflorescence stems, flowers and siliques. Highest expression in roots and rosette leaves. Very low expression in flowers. {ECO:0000269|PubMed:9862967}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds specifically to bipartite recognition sequences composed of two unrelated motifs, 5'-CAACA-3' and 5'-CACCTG-3'. May function as negative regulator of plant growth and development. {ECO:0000269|PubMed:15040885, ECO:0000269|PubMed:9862967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
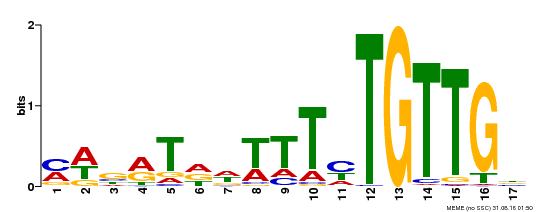
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009148531.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by brassinosteroid and zeatin. {ECO:0000269|PubMed:15040885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353075 | 0.0 | AK353075.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-20-J24. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009148531.1 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| Swissprot | Q9ZWM9 | 0.0 | RAV1_ARATH; AP2/ERF and B3 domain-containing transcription factor RAV1 | ||||
| TrEMBL | A0A3P5YPI2 | 0.0 | A0A3P5YPI2_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4DTH8 | 0.0 | M4DTH8_BRARP; Uncharacterized protein | ||||
| STRING | Bra019821.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 0.0 | related to ABI3/VP1 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



