 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009143231.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 407aa MW: 45728.6 Da PI: 9.6688 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 97.8 | 7.7e-31 | 80 | 133 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtpeLH rF++ave+ GG+++AtPk +l+lm+vkgL ++hvkSHLQ+YR+
XP_009143231.1 80 PRLRWTPELHLRFLQAVEKFGGPDRATPKLVLQLMNVKGLCIAHVKSHLQMYRS 133
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.223 | 76 | 136 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-29 | 76 | 134 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.29E-15 | 77 | 135 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-23 | 80 | 135 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.0E-10 | 81 | 132 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 407 aa Download sequence Send to blast |
MRSSSQSSEN SKTCPSNNFK ETTKKEDNKD DENKDEEARS VDQIPSNNSY VEESGSHHHN 60 NDQIKKNGGS VRPYNRSKTP RLRWTPELHL RFLQAVEKFG GPDRATPKLV LQLMNVKGLC 120 IAHVKSHLQM YRSKKTDDPN QGNQGFSFEH GAGYTYNLGQ LPMLQSFDQR PSTSLGYDGG 180 SWTDRRRQVY RSPWRGLTAR NSTRTRPTLF SSQSGERFRG VSNSILDDKN KTISFRINSH 240 EAAQASNGVG ESVPSIHRSF LEGTKTLDKS WGQSLSSNPN YSTASGPQDH IATTLSSNQR 300 ENPRVAEETE NVLKRKRLLL SDDCNKSVQD LDLSLSLKVH RTHNNLGDCL LEDEEKKLDD 360 HEDIKGLSLS LASSSSSKLG RDIRKEVQND TKKRKISVLA SPLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-16 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-16 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-16 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-16 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
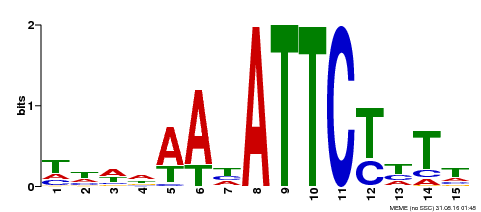
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009143231.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009143231.1 | 0.0 | PREDICTED: uncharacterized protein LOC103866965 isoform X2 | ||||
| TrEMBL | M4CL87 | 0.0 | M4CL87_BRARP; Uncharacterized protein | ||||
| STRING | Bra004973.1-P | 0.0 | (Brassica rapa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 0.0 | G2-like family protein | ||||



