 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009125296.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 566aa MW: 63656.9 Da PI: 6.8958 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.6 | 7.6e-14 | 265 | 324 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +++Aa+a++ a++k+++
XP_009125296.1 265 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEDKAARAYDLAALKYWN 324
57*******************666664478446**********99*************97 PP
| |||||||
| 2 | AP2 | 49.7 | 9.3e-16 | 367 | 418 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
XP_009125296.1 367 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 418
57****************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.72E-20 | 265 | 334 | No hit | No description |
| SuperFamily | SSF54171 | 3.66E-16 | 265 | 333 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 8.4E-11 | 265 | 324 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.1E-25 | 266 | 338 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.5E-14 | 266 | 332 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 18.545 | 266 | 332 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.0E-6 | 267 | 278 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.59E-11 | 367 | 426 | No hit | No description |
| Pfam | PF00847 | 3.9E-10 | 367 | 418 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.05E-17 | 367 | 427 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 19.111 | 368 | 426 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.3E-30 | 368 | 432 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.3E-18 | 368 | 426 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.0E-6 | 408 | 428 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 566 aa Download sequence Send to blast |
MAPMTNWLTF SLSPMDMLRS SDQSQFVSYD ASSAASSSPY LLDNFYGWTN QKPQEFFKDE 60 AQIAASMADS TILTTFVDPQ THSHNHIPKL EDFLGEVRYS DNSQTETQDS SSLTHIYDPR 120 HHQNQNQTGF YSDHNHEFKT MAGFQTAFST NSGSEVEDSA SIGRTHLAGE YLGHVVESSG 180 GPELGFHGGA NNGGALSLGV NVNNSNHRTS DDHTQITEHH YRGNNNGERI NNEKMVSEKE 240 KPVVAVETSD CSNKKIADTF GQRTSIYRGV TRHRWTGRYE AHLWDNSCRR EGQARKGRQV 300 YLGGYDKEDK AARAYDLAAL KYWNATATTN FPITNYSKEL EEMKHMTKQE FIASLRRKSS 360 GFSRGASIYR GVTRHHQQGR WQARIGRVAG NKDLYLGTFA TEEEAAEAYD IAAIKFRGIN 420 AVTNFEMNRY DIEAIMKSAL PIGGAAKRLK LSLEAAEQKP ILGHQHQLHH FQQQQQQQIQ 480 SSPNHSSINF AQSQMIPCGI PFEAAALYHH QQQQQQQQQQ NFFQHFPANV QASDSTGSNN 540 NSNVQGSMGL MTPNQAEFFL WPNQSY |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.29870 | 0.0 | root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Detected in inflorescence and youg floral mersitems, and in stem procambial cells. In floral mersitems, mostly expressed in the central dome. Disappears progressively from sepal primordia, but accumulates in second, third and fourth whorl organ primordia. Later, confined to occasional patches in stamens and in petal before disparearing progressively from flowers. {ECO:0000269|PubMed:15988559}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, seedlings, hypocotyl, inflorescence, siliques, and pistils. Also detected at low levels in leaves. {ECO:0000269|PubMed:15988559, ECO:0000269|PubMed:16307362}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
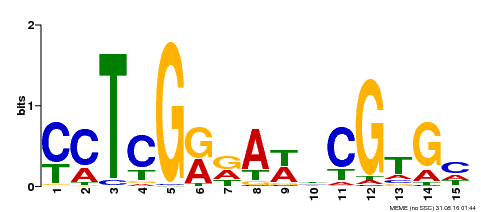
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009125296.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY585683 | 0.0 | AY585683.1 Arabidopsis thaliana clone at5g10510 AP2/EREBP transcription factor mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009125296.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A398A6V8 | 0.0 | A0A398A6V8_BRACM; Uncharacterized protein | ||||
| STRING | Bra028584.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3333 | 27 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



