 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009124485.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 889aa MW: 99068.5 Da PI: 6.2445 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 134.6 | 3.1e-42 | 134 | 211 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+Ceadls++k+yhrrhkvCe+hska+++lv+g++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
XP_009124485.1 134 VCQVESCEADLSKVKDYHRRHKVCEMHSKATSALVAGIMQRFCQQCSRFHVLEEFDEGKRSCRRRLAGHNKRRRKTNP 211
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.0E-34 | 127 | 196 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.488 | 132 | 209 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 4.97E-39 | 134 | 213 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 9.3E-31 | 135 | 208 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 2.4E-7 | 675 | 781 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.71E-7 | 677 | 778 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.49E-8 | 680 | 778 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 889 aa Download sequence Send to blast |
MEARVEGEGQ HFYGYRGKRS VEWDLNDWKW DGDLFIATPL NPGVSQTMGR QFFPLGNSSN 60 SSSSCSEEGN GNITREVEKR RRAAAAATGE DTDNNNNNGG LSLKLGENGY DLNGEREGKK 120 TKLGGGSGTN HRSVCQVESC EADLSKVKDY HRRHKVCEMH SKATSALVAG IMQRFCQQCS 180 RFHVLEEFDE GKRSCRRRLA GHNKRRRKTN PEPAANGNPL SDDNQSSNYL LICLLKILSN 240 MHSNGSSDHQ DLMPHLLKSL VSHAGEQLGK NLVELLLQGG SGLLAAPQED SKQAPEIPRQ 300 ELYANGNRSE KLQTKVNDFD LNDIYIDSDD GTDLERSSPP TTTTNPATSS PDYPSWIHQT 360 SRNSDSASDQ SPSSSSEDAQ MRTGRIVFKL FGKEPNDFPP VLRGQILDWL SHTPTDIESY 420 IRPGCIVLTI YLRQAETAWE ELSDDMGFSL SKLLDLSDDP LWTSGWIYVR MQNQYAFVFD 480 GQVVVDTSLP LRSYDHSHII SVRPLAVAAT GKAQFTVKGI NLRRPGTRLL CAVGGKYLIQ 540 ENDDLKESNE CVSFSCDLPI TSGRGFMEIE DQGGLSSSFF PFIVVEEDDV CSEIRILETT 600 LEFTDTDSAK LAMEFIHELG WLLRRSKLGV FSLARFKWLI EFSMDREWCA VIRKLLNMFF 660 EGAVGDSSSD AAALSELCLL HRAVRKNSKP MVEMLLRYVV PNQQRIHSLF RPDAAGPAGL 720 TPLHIAAGKD GSEDVLDALT EDPSMVGIEA WRTSKDTTGF TPEDYARLRG HFSYIHLIQR 780 KINKKSATED HVVVNIPASS ISDREQKETK SGSSALEITH GNNKLQCKLC DHKLVYGTAR 840 RSVAYRPAML SMVAIAAVCV CVALLFKSCP EVLYVFQPFR WELLDYGTR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-33 | 125 | 208 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.16500 | 0.0 | root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
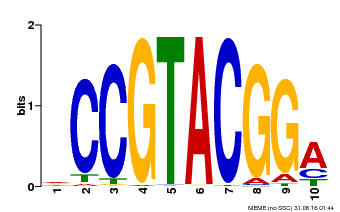
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009124485.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009124485.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | M4FIV8 | 0.0 | M4FIV8_BRARP; Uncharacterized protein | ||||
| STRING | Bra041037.1-P | 0.0 | (Brassica rapa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



