 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009122730.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 506aa MW: 55681.3 Da PI: 10.1299 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.2 | 1.4e-05 | 82 | 104 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++C+k F r nL+ H r H
XP_009122730.1 82 FVCEICNKGFQRDQNLQLHRRGH 104
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 15 | 7e-05 | 159 | 181 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k + +s+ k H +t+
XP_009122730.1 159 FKCEKCSKKYAVQSDWKAHAKTC 181
89******************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.19E-7 | 81 | 104 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-5 | 81 | 104 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF12171 | 8.1E-5 | 82 | 104 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE profile | PS50157 | 10.949 | 82 | 104 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.017 | 82 | 104 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 84 | 104 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 73 | 124 | 154 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 147 | 180 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.19E-7 | 154 | 179 | No hit | No description |
| SMART | SM00355 | 92 | 159 | 179 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0045604 | Biological Process | regulation of epidermal cell differentiation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 506 aa Download sequence Send to blast |
MQMIPGDPFS ISSSMGGFVH QEQHLHHLQQ HIQDINPNSN PNPNAKPNSS SAKKKRNQPG 60 TPDPDADVIA LSPTTLMATN RFVCEICNKG FQRDQNLQLH RRGHNLPWKL KQRSNKEVIK 120 KKVYICPIKT CVHHDSSRAL GDLTGIKKHY SRKHGEKKFK CEKCSKKYAV QSDWKAHAKT 180 CGTREYKCDC GTLFSRKDSF ITHRAFCDAL TEEGARMSSL SNNSAISTTT MTTNLNFGND 240 SHVMNNSNLP HGFVHRGVHH PDINAAISQF GLGFGHDLTG MHPQGLSEMV QMTSSGNHHL 300 FPLSSSSVLP DFSGHHHNQQ FQIPTTSTTS QQTSASLPSL HQQHQTLKDS SFSPLFSSSS 360 ETKQNKPLSP MSATALLQKA AQMGSTRSNS TTAPSIFAGP TMTSSSSAAS PPPRSSSPMM 420 IQQQLNNNFN TNVPREIHNL APPSSGVTTS SVDNNHRFQS NRSGITPVQQ MGLTRDFLGV 480 SNEHHPHQTG HRPFLPQELA RFAPSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 4e-42 | 155 | 224 | 3 | 72 | Zinc finger protein JACKDAW |
| 5b3h_F | 4e-42 | 155 | 224 | 3 | 72 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Start to accumulate during the 16- to 32-cell stage of embryogenesis. {ECO:0000269|PubMed:17785527}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the quiescent center, the ground tissue stem cells and to a lesser extent in mature cortex and endodermis cells. {ECO:0000269|PubMed:17785527}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that, together with BIB, regulates tissue boundaries and asymmetric cell division by a rapid up-regulation of 'SCARECROW' (SCR), thus controlling the nuclear localization of 'SHORT-ROOT' (SHR) and restricting its action (PubMed:17785527, PubMed:25829440). Binds DNA via its zinc fingers (PubMed:28211915, PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). Confines CYCD6 expression to the cortex-endodermis initial/daughter (CEI/CEID) tissues (PubMed:25829440). Required for radial patterning and stem cell maintenance (PubMed:17785527). Counteracted by 'MAGPIE' (MGP) (PubMed:17785527). Binds to the SCR and MGP promoter sequences (PubMed:21935722). Controls position-dependent signals that regulate epidermal-cell-type patterning (PubMed:20356954). {ECO:0000269|PubMed:17785527, ECO:0000269|PubMed:20356954, ECO:0000269|PubMed:21935722, ECO:0000269|PubMed:24821766, ECO:0000269|PubMed:25829440, ECO:0000269|PubMed:28211915}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00485 | DAP | Transfer from AT5G03150 | Download |
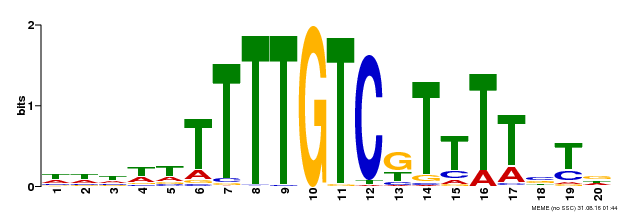
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009122730.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by SCR and SHR. {ECO:0000269|PubMed:21935722}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ621494 | 0.0 | AJ621494.1 Arabidopsis thaliana mRNA for ID1-like zinc finger protein 3 (idz3 gene). | |||
| GenBank | AJ630493 | 0.0 | AJ630493.1 Arabidopsis thaliana mRNA for hypothetical protein, clone At5g03150. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009122730.1 | 0.0 | PREDICTED: zinc finger protein JACKDAW isoform X2 | ||||
| Swissprot | Q700D2 | 0.0 | IDD10_ARATH; Zinc finger protein JACKDAW | ||||
| TrEMBL | A0A397XP91 | 0.0 | A0A397XP91_BRACM; Uncharacterized protein | ||||
| STRING | Bra009549.1-P | 0.0 | (Brassica rapa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G03150.1 | 0.0 | C2H2-like zinc finger protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



