 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009122177.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 270aa MW: 30609.4 Da PI: 8.3935 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.5 | 6.5e-08 | 30 | 75 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqk 46
WT+eE +++ +a + + + +W ++a++++ g+t ++ + k
XP_009122177.1 30 TWTKEENKKFERALAIYADDtpdRWFKVAAMIP-GKTISDVMTQYSK 75
7*****************99*************.****999888866 PP
| |||||||
| 2 | Myb_DNA-binding | 40 | 9e-13 | 130 | 174 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++G+g+W+ I+r + +t+ q+ s+ qky
XP_009122177.1 130 PWTEEEHRRFLLGLLKYGKGDWRNISRNFVGSKTPTQVASHAQKY 174
8****************************999************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.121 | 26 | 79 | IPR017884 | SANT domain |
| SMART | SM00717 | 1.7E-7 | 27 | 79 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.15E-10 | 29 | 83 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.7E-7 | 29 | 75 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.52E-7 | 30 | 76 | No hit | No description |
| PROSITE profile | PS51294 | 18.108 | 123 | 179 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.0E-15 | 125 | 178 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-11 | 127 | 179 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-16 | 127 | 178 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 9.9E-11 | 127 | 177 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.75E-10 | 130 | 174 | No hit | No description |
| Pfam | PF00249 | 3.8E-10 | 130 | 174 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 270 aa Download sequence Send to blast |
METLHPLLSH VPTSDHRFVV QEMMCLQTST WTKEENKKFE RALAIYADDT PDRWFKVAAM 60 IPGKTISDVM TQYSKLEEDL FDIEAGLVPI PGYPSAGFDQ LVSPYDHDLY RKRPNGGARG 120 FDQDRRKGVP WTEEEHRRFL LGLLKYGKGD WRNISRNFVG SKTPTQVASH AQKYYQRQLS 180 GAKDKRRPSI HDITTVNLLN TNITRPSSDH DRFLQQPDES KLGFTDKGNA EEGVMFLGQN 240 LSSVLSPYEP AFKFTGTNVY GGGAYGLARS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 9e-16 | 31 | 97 | 11 | 77 | RADIALIS |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.26810 | 0.0 | leaf| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Detected in the apical inflorescence meristem, in bract primordia arising in its periphery and in floral meristems produced in the axils of bracts (stages 0-3). From stage 3 to stage 8, detected in all floral organs irrespective of their dorsoventral positions. From stage 9, barely detectable in bracts, sepals, and stamens. In the corolla, however, expression was maintained and enhanced in some regions. Within ventral and lateral petals at stage 9, asymmetric pattern of expression with high levels of transcripts in the inner epidermis of the furrow and very reduced levels in the remaining cell layers. In the dorsal petals, from stage 9 onward, detected but with a more even distribution across cell layers than in the ventral petal. {ECO:0000269|PubMed:11937495}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00489 | DAP | Transfer from AT5G05790 | Download |
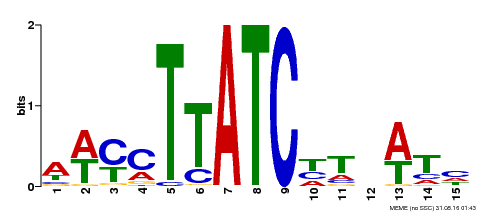
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009122177.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC189201 | 0.0 | AC189201.1 Brassica rapa subsp. pekinensis cultivar Inbred line 'Chiifu' clone KBrB005N03, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009122177.1 | 0.0 | PREDICTED: transcription factor DIVARICATA | ||||
| Refseq | XP_013715256.1 | 0.0 | transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 8e-71 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A397XQG4 | 0.0 | A0A397XQG4_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4CY37 | 0.0 | M4CY37_BRARP; Uncharacterized protein | ||||
| STRING | Bra009134.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4340 | 26 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G05790.1 | 1e-165 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



