 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bathy15g00870 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Bathycoccus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 581aa MW: 61318.8 Da PI: 9.8135 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 93.1 | 2.2e-29 | 341 | 394 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av+qL G++kA+Pk+il+lm+v+gLt+e+v+SHLQkYRl
Bathy15g00870 341 KPRVVWSAELHQQFVNAVNQL-GIDKAVPKRILDLMNVQGLTRENVASHLQKYRL 394
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 83 | 9.4e-28 | 14 | 122 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTE CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGa 94
vl+vdD+ l +++++++l+k gy v++++++ ale+l++k+ +D++l D+ mp+mdG++ll++i+ ++p+++++ +++++ +l+ + Ga
Bathy15g00870 14 VLVVDDDVLCLKIVTKMLQKCGY-TVTSTTSSVTALEYLRTKKeeFDIVLSDVHMPDMDGFKLLEQIALDI-DIPVLMMSVNADQSVVLRGIIHGA 107
89*********************.***************777777**********************9877.9*********************** PP
SEEEESS--HHHHHH CS
Response_reg 95 kdflsKpfdpeelvk 109
d+l Kp+ + el +
Bathy15g00870 108 VDYLLKPVRIHELKN 122
**********99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF52172 | 6.69E-35 | 10 | 129 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 2.8E-39 | 11 | 135 | No hit | No description |
| SMART | SM00448 | 9.0E-32 | 12 | 124 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.443 | 13 | 128 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.3E-24 | 14 | 122 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 2.51E-25 | 15 | 127 | No hit | No description |
| PROSITE profile | PS51294 | 12.084 | 338 | 397 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.3E-31 | 339 | 399 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.55E-20 | 339 | 398 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-25 | 341 | 394 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.4E-8 | 343 | 393 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009736 | Biological Process | cytokinin-activated signaling pathway | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 581 aa Download sequence Send to blast |
MSGRSTPTIP NLRVLVVDDD VLCLKIVTKM LQKCGYTVTS TTSSVTALEY LRTKKEEFDI 60 VLSDVHMPDM DGFKLLEQIA LDIDIPVLMM SVNADQSVVL RGIIHGAVDY LLKPVRIHEL 120 KNIWQHVVRK NGPDSLQRSG SMSMSPARSG RGSPEKDSGM NATSNGTHMN KKLRTIAAAN 180 NAVSTDASAA AKGNKKPSTV EGVAQLQQQQ QQQQQTTTSG NDNNNNNNNN STAPVPLPGG 240 AGPGQQQQQN QINVQGHLGG GHPPLQSTHQ QQQQQQHVDG SASAIPSARR PPPMGYLDPN 300 NMMSGNPTTA GGVDMNGMPM QGNGAPGAGQ SGGSSGGGSK KPRVVWSAEL HQQFVNAVNQ 360 LGIDKAVPKR ILDLMNVQGL TRENVASHLQ KYRLYLKRLQ GGPNNPSGPG FLSNKIAGGA 420 TGASKPSGSG KSKNNASKVS VPGGSYQFHP GVGAPPQQWN QGMPPQQHQQ QQQQQQQQAP 480 PHMVAAGGPG GMPPQQHPPQ YSTFPHPQGV ITPGPPPPGT VRYGYGPVPI DASSFTPIPL 540 DFDDTGLGLG IGTSKGSFGS KSTDDMLSMF LKDGDPESIL * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-22 | 338 | 399 | 2 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
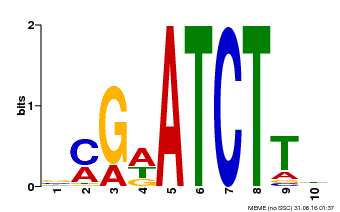
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FO082264 | 0.0 | FO082264.1 Bathycoccus prasinos genomic : Chromosome_15. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007508971.1 | 0.0 | type-b response regulator | ||||
| TrEMBL | K8EP92 | 0.0 | K8EP92_9CHLO; Two-component response regulator | ||||
| STRING | XP_007508971.1 | 0.0 | (Bathycoccus prasinos) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP1220 | 15 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-36 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 19011598 |



