 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bathy10g00280 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Bathycoccus
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 672aa MW: 72448.6 Da PI: 9.4697 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 47.8 | 2.9e-15 | 345 | 381 | 3 | 40 |
TCR 3 kkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
+k+CnCkkskClk+YC+Cfaag++C + C+C++C+N+e
Bathy10g00280 345 SKKCNCKKSKCLKLYCDCFAAGVFCRD-CSCQSCSNTE 381
79*************************.********97 PP
| |||||||
| 2 | TCR | 50.5 | 4e-16 | 415 | 454 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
k+ kgC+Ckks ClkkYCeCf+a++ C++ CkCe+CkN++
Bathy10g00280 415 KHAKGCHCKKSACLKKYCECFQANVRCQDYCKCEGCKNTT 454
589***********************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 1.3E-14 | 343 | 383 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 31.419 | 344 | 455 | IPR005172 | CRC domain |
| Pfam | PF03638 | 8.0E-12 | 346 | 381 | IPR005172 | CRC domain |
| SMART | SM01114 | 7.5E-14 | 415 | 456 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 4.1E-12 | 418 | 453 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 672 aa Download sequence Send to blast |
MEVSFVTVFF EVHVSRKMTP SFFTILTSHA TYKMGISNMD DIMKTPERPK KNGNGDIDPS 60 IINSPFMNFV SNLSPIRFGA SSLGGNNTVV PICFSSPSAF KSPPRKRSED NGNHNHNNTV 120 SSLANDMGPI PDSLMVPFAH EKHQHEDADD QDDFVGGIGG TMGAITLEDE LLQLDNKFNN 180 KVNNNKHNNS GGKGDKTTAK KSGGDQKKSA KKNGGKNNNA SVSPPTGSSK RQSTTGASLE 240 WNSGFSKTIR GKAADSPRDH PVGTENRKAG ATPMASTKKS SRRDSLTSSN DEHITPPVGK 300 SKSDGGSKVK NTRDLQTPGS NTKLTFADKL DQSKLDEGID FDGASKKCNC KKSKCLKLYC 360 DCFAAGVFCR DCSCQSCSNT EGDLEVVRQT RYQIESRNPN AFANKIVDDD SVDAKHAKGC 420 HCKKSACLKK YCECFQANVR CQDYCKCEGC KNTTDGANPS QNGGAATATK TTITTTTTTT 480 AVREDEGKIL TNTNEAKKKV EGLKKSIVGS VRDSDTSFIL NSPAREAARA AQRMLADDVF 540 MDDMSFVNYG HGNLASPLRT LLHSETGLSP MFQRVNDDKG ATATNSRQCK KENNDVNANN 600 KSNSVGNNTK ASSKTRGAAG RFSFSKTNSA MNTRRKAPVP LFTDDTNNVE EKTPARTTRS 660 RGSPKLVTPS V* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 6e-19 | 346 | 455 | 12 | 123 | Protein lin-54 homolog |
| 5fd3_B | 6e-19 | 346 | 455 | 12 | 123 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
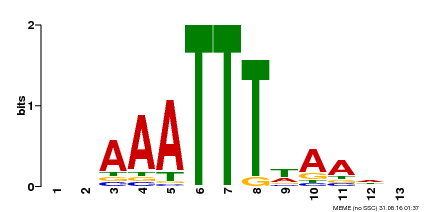
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FO082269 | 0.0 | FO082269.1 Bathycoccus prasinos genomic : Chromosome_10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007510801.1 | 0.0 | predicted protein | ||||
| TrEMBL | K8EK20 | 0.0 | K8EK20_9CHLO; Uncharacterized protein | ||||
| STRING | XP_007510801.1 | 0.0 | (Bathycoccus prasinos) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP323 | 15 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14770.1 | 5e-25 | TESMIN/TSO1-like CXC 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 19013105 |



