 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013635980.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 871aa MW: 96802.1 Da PI: 6.1232 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 134.8 | 2.7e-42 | 114 | 191 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+Ceadls++k+yhrrhkvCe+hska+++lv g++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
XP_013635980.1 114 VCQVESCEADLSKVKDYHRRHKVCEMHSKATSALVGGIMQRFCQQCSRFHVLEEFDEGKRSCRRRLAGHNKRRRKTNP 191
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.5E-34 | 107 | 176 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.513 | 112 | 189 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.79E-39 | 114 | 193 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.3E-31 | 115 | 188 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 3.8E-7 | 656 | 763 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.73E-7 | 659 | 763 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.07E-8 | 662 | 760 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 871 aa Download sequence Send to blast |
MDVGKRSVEW DLNDWKWDGD LFIATPLNPG VSQTMGRQFF PLGNSSNTSS SCSDEGNDNT 60 REVEKRRRAA ATGEDGGGGG LSLKLGENGY DLNGEREGKK TKLGGGSGTN HRSVCQVESC 120 EADLSKVKDY HRRHKVCEMH SKATSALVGG IMQRFCQQCS RFHVLEEFDE GKRSCRRRLA 180 GHNKRRRKTN PEPAANGNPL SDDNQSSNYL LICLLKILSN MHSNGSSGDQ DLMPHLLKSL 240 VSHAGEQLGK NLVELLLQGG SGLLAAPQED SKQAPELPRQ ELYASGNRSE KLQTKVNDFD 300 LNDIYIDSDD GGTDLERSSP PTTTTNPATS SPDYPSWIHQ TSRNSDSASD QSPSSSSEDA 360 QMRTGRIVFK LFGKEPNDFP PVLRGQILDW LSHTPTDIES YIRPGCIVLT IYLRQAETAW 420 EELSDDMGFS LSKLLDLSDD PLWTSGWIYV RMQNQYAFVF DGQVVVDSSL PLRSHDYSHI 480 ISVRPLAVAA ATGKAQFTVK GINLRRPGTR LLCAVEGKYL IQENDDLKES NECVSFSCDL 540 PITSGRGFME IEDQGGLSSS FFPFIVVEED DVCSEIRILE TTLEFTDTDS AKLAMEFIHE 600 LGWLLRRSKL GVFSLARFKW LIEFSMDREW CAVIRKLLNM LFEGAVGDTS SDAAALSELC 660 LLHRAVRKNS KPMVEMLLRY VVPNQQRIHS LFRPDSAGPA GLTPLHIAAG KDGSEDVLDA 720 LTEDPSMVGI EAWRTSRDTT GFTPEDYARL RGHFSYIHLI QRKINKKSAT EDHVVVNIPA 780 SSISDREQKE TKSGSSALEI THGNHKLQCK LCDHKLVYGT ARRSVAYRPA MLSMVAIAAV 840 CVCVALLFKS CPEVLYVFQP FRWELLDYGT R |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-32 | 105 | 188 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
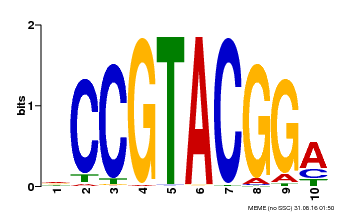
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013635980.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013635980.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 isoform X4 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A0D3BNN1 | 0.0 | A0A0D3BNN1_BRAOL; Uncharacterized protein | ||||
| STRING | Bo4g005730.1 | 0.0 | (Brassica oleracea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



