 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013635978.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 888aa MW: 98798.3 Da PI: 6.1036 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 134.8 | 2.8e-42 | 131 | 208 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+Ceadls++k+yhrrhkvCe+hska+++lv g++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
XP_013635978.1 131 VCQVESCEADLSKVKDYHRRHKVCEMHSKATSALVGGIMQRFCQQCSRFHVLEEFDEGKRSCRRRLAGHNKRRRKTNP 208
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.7E-34 | 124 | 193 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.513 | 129 | 206 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.92E-39 | 131 | 210 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.4E-31 | 132 | 205 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 3.9E-7 | 673 | 780 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.73E-7 | 676 | 780 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.99E-8 | 679 | 777 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 888 aa Download sequence Send to blast |
MEARIEGEGQ HFYGYRAMDV GKRSVEWDLN DWKWDGDLFI ATPLNPGVSQ TMGRQFFPLG 60 NSSNTSSSCS DEGNDNTREV EKRRRAAATG EDGGGGGLSL KLGENGYDLN GEREGKKTKL 120 GGGSGTNHRS VCQVESCEAD LSKVKDYHRR HKVCEMHSKA TSALVGGIMQ RFCQQCSRFH 180 VLEEFDEGKR SCRRRLAGHN KRRRKTNPEP AANGNPLSDD NQSSNYLLIC LLKILSNMHS 240 NGSSGDQDLM PHLLKSLVSH AGEQLGKNLV ELLLQGGSGL LAAPQEDSKQ APELPRQELY 300 ASGNRSEKLQ TKVNDFDLND IYIDSDDGGT DLERSSPPTT TTNPATSSPD YPSWIHQTSR 360 NSDSASDQSP SSSSEDAQMR TGRIVFKLFG KEPNDFPPVL RGQILDWLSH TPTDIESYIR 420 PGCIVLTIYL RQAETAWEEL SDDMGFSLSK LLDLSDDPLW TSGWIYVRMQ NQYAFVFDGQ 480 VVVDSSLPLR SHDYSHIISV RPLAVAAATG KAQFTVKGIN LRRPGTRLLC AVEGKYLIQE 540 NDDLKESNEC VSFSCDLPIT SGRGFMEIED QGGLSSSFFP FIVVEEDDVC SEIRILETTL 600 EFTDTDSAKL AMEFIHELGW LLRRSKLGVF SLARFKWLIE FSMDREWCAV IRKLLNMLFE 660 GAVGDTSSDA AALSELCLLH RAVRKNSKPM VEMLLRYVVP NQQRIHSLFR PDSAGPAGLT 720 PLHIAAGKDG SEDVLDALTE DPSMVGIEAW RTSRDTTGFT PEDYARLRGH FSYIHLIQRK 780 INKKSATEDH VVVNIPASSI SDREQKETKS GSSALEITHG NHKLQCKLCD HKLVYGTARR 840 SVAYRPAMLS MVAIAAVCVC VALLFKSCPE VLYVFQPFRW ELLDYGTR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-32 | 122 | 205 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
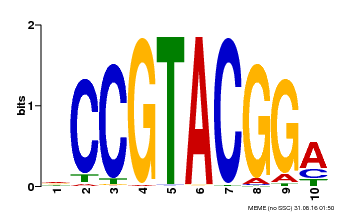
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013635978.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013635978.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A0D3BNN1 | 0.0 | A0A0D3BNN1_BRAOL; Uncharacterized protein | ||||
| STRING | Bo4g005730.1 | 0.0 | (Brassica oleracea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



