 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013623193.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 427aa MW: 47833.3 Da PI: 5.9289 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.5 | 2.3e-31 | 225 | 279 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr+rWtpeLHe+F++ v++L G+ekAtPk++l+lm+v+gLt++hvkSHLQkYRl
XP_013623193.1 225 KPRMRWTPELHESFLKSVDKLEGPEKATPKAVLKLMNVEGLTIYHVKSHLQKYRL 279
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 9.961 | 222 | 282 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-29 | 223 | 280 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.33E-16 | 224 | 280 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.4E-24 | 225 | 280 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.0E-8 | 227 | 278 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 2.5E-22 | 315 | 362 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 427 aa Download sequence Send to blast |
MNSHRLVSTA QDECNKGLHQ PCSSSLSPVH NFLNLQPENR NSPFIRSQSP DSPWPKNSPQ 60 GTFSRSSTFC TNLYISSSST SDAHKHLGNT LPFLPDPSTN IQPPSGVESA RSPSIFSEDM 120 SNPVDGDNTL VKDFFNLSGD ACSDRAFHDL DCSNDSYCLS DQMELQFLSD ELELAITDRS 180 ETPRLDEIYE TPLASSSNPV TGMSQSQRCV TGAVSTGSAA ASNHKPRMRW TPELHESFLK 240 SVDKLEGPEK ATPKAVLKLM NVEGLTIYHV KSHLQKYRLA KHIPEKKEEK KNVNTEEKKL 300 ALSNSEAVER KKGAMQLTEA LRMQMEVQKQ LHEQLEVQRV LQLRIEEHAK YLEKMLEEQR 360 KTGMLMSSSS SSSQTLLSDC QNMSKTEVSS LSQPKNIASE TEDDKCESPQ KRRRVENNTE 420 SEDPPER |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-23 | 225 | 282 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 410 | 414 | KRRRV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
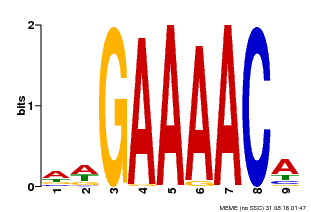
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013623193.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013623193.1 | 0.0 | PREDICTED: protein PHR1-LIKE 1-like isoform X1 | ||||
| Swissprot | Q949U2 | 0.0 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | A0A0D3BB17 | 0.0 | A0A0D3BB17_BRAOL; Uncharacterized protein | ||||
| TrEMBL | A0A3P6BBX1 | 0.0 | A0A3P6BBX1_BRAOL; Uncharacterized protein | ||||
| STRING | Bo3g065290.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6786 | 27 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 0.0 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



