 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013612299.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 791aa MW: 86540.5 Da PI: 6.0523 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.7e-21 | 124 | 179 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
XP_013612299.1 124 KKRYHRHTPQQIQELESMFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 179
688999***********************************************999 PP
| |||||||
| 2 | START | 206.3 | 1.2e-64 | 311 | 531 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevi 85
ela++a++el+k+a+++ep+Wvks +++n +e++++f+++k +ea+r+sg+v+ ++ lve+l+d+ +W+e+++ + +t++vi
XP_013612299.1 311 ELALTAMDELLKLAQSDEPLWVKSLdgerDELNHEEYMRTFSSTKPngLVTEASRTSGMVIINSLSLVETLMDSD-RWTEMFPcivaRSTTTDVI 404
5899*************************************99999999**************************.******************* PP
CTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE- CS
START 86 ssg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwvehvdl 172
s g ga+qlm+aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++++ s v+R +lpSg++++++sng+skvtwveh+++
XP_013612299.1 405 SGGmagtrnGAIQLMNAELQVLSPLVPvRNVNFLRFCKQHAEGVWAVVDVSIDTVRENSGgSTVVIR--RLPSGCVVQDMSNGYSKVTWVEHAEY 497
***********************************************************76666666..************************** PP
-SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 173 kgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++h l+r+l++sgl +g ++wvatlqrqce+
XP_013612299.1 498 DENQIHHLYRPLLRSGLGFGSQRWVATLQRQCEC 531
********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-22 | 111 | 181 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.2E-20 | 111 | 181 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.281 | 121 | 181 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 122 | 185 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.46E-18 | 124 | 181 | No hit | No description |
| Pfam | PF00046 | 2.0E-18 | 124 | 179 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 156 | 179 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.819 | 302 | 534 | IPR002913 | START domain |
| CDD | cd08875 | 1.30E-115 | 306 | 530 | No hit | No description |
| SuperFamily | SSF55961 | 1.92E-31 | 306 | 531 | No hit | No description |
| Pfam | PF01852 | 9.4E-58 | 311 | 531 | IPR002913 | START domain |
| SMART | SM00234 | 3.2E-41 | 311 | 531 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.24E-21 | 551 | 764 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 791 aa Download sequence Send to blast |
MNFGSLFDNT TGGVPTGLAY GNHTTATTVI PGGAMSHIAA ASLFSPPINT KSIYSSSGLS 60 LTLEQPERGT SMRNNNGAGG DIFDGSGKNR RSREEEHESR SGSDNVEGIS GEDQDADDNK 120 PPKKKRYHRH TPQQIQELES MFKECPHPDE KQRLELSKRL CLETRQVKFW FQNRRTQMKT 180 QLERHENALL RQENDKLRAE NMSIREAMRN PTCNICGGPA MLGDVSIEEH HLRIENARLK 240 DELDRVFNLT GKFLGHHHNN HTSSLELGVG TNNNGGNFAF PPDFNGGGGC LPPPQQATVI 300 NGVDQRSVLL ELALTAMDEL LKLAQSDEPL WVKSLDGERD ELNHEEYMRT FSSTKPNGLV 360 TEASRTSGMV IINSLSLVET LMDSDRWTEM FPCIVARSTT TDVISGGMAG TRNGAIQLMN 420 AELQVLSPLV PVRNVNFLRF CKQHAEGVWA VVDVSIDTVR ENSGGSTVVI RRLPSGCVVQ 480 DMSNGYSKVT WVEHAEYDEN QIHHLYRPLL RSGLGFGSQR WVATLQRQCE CLAILMSSSV 540 TSHDDTSISP GGRKSMLKLA QRMTINFCSG ISAPSVHSWS KLTVGNVDPD VRIMTRKSVD 600 DPSEAPGIVL SAATSVWLPA SPQRLFDFLR NERMRCEWDI LSNGGPMQEM AHIAKGQDQG 660 NSVSLLRSNP MNANQSSMLI LQETCIDASG ALVVYAPVDI PAMHVVMNGG DSSYVALLPS 720 GFAVLPDGGF DGVSGDGEQR PVGGGSLLTV AFQILVNNLP TAKLTVESVE TVNSLISCTV 780 QKIRTALQCE N |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bol.2235 | 6e-86 | leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
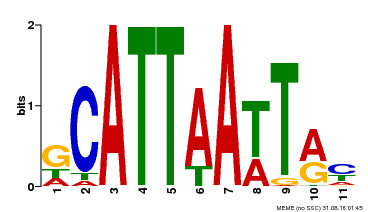
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013612299.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013612299.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A0D3DZP0 | 0.0 | A0A0D3DZP0_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g002480.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



