 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013610197.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 359aa MW: 39162.6 Da PI: 8.3569 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.9 | 1e-41 | 105 | 182 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+C v+gC++d+s+++eyh+rhkvCevhsk+pvv+++g++qrfCqqCsrfh+l+efDe+krsCr+rL++hn+rrrk+q+
XP_013610197.1 105 ICLVDGCDSDFSNCREYHKRHKVCEVHSKTPVVTINGHKQRFCQQCSRFHALEEFDEGKRSCRKRLDGHNRRRRKPQP 182
5**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.6E-33 | 100 | 167 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.06 | 103 | 180 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.31E-39 | 104 | 186 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.4E-30 | 106 | 179 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 359 aa Download sequence Send to blast |
MDWNFKLSSG YFSEFDQESV PDIAPIDGSI SFGGSSSPRL QPEGDFSFDL KLGRNIGNSS 60 SSAYGNTEQV PSLCKWKESK MAKPRSSGSS KRKRGNAIGN SQIPICLVDG CDSDFSNCRE 120 YHKRHKVCEV HSKTPVVTIN GHKQRFCQQC SRFHALEEFD EGKRSCRKRL DGHNRRRRKP 180 QPDHISRPAN FFAGFQGSKM LEFSSSSYVF PTTSVASPSW GNGLVSVAMA NGSSNGQNQG 240 VSGSSPAKTG IMFPISSSSS SRRKLLPSLQ EQESSRTAPL CERMTSCIHD SDCALSLLSS 300 SSSSVPHLLQ PPLSLYQEAA ETGFYGLGLF ENATAVSDGS VISSIEAVAL PQTFPFHWE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-29 | 99 | 179 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 162 | 179 | KRSCRKRLDGHNRRRRKP |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Weak increase of expression during floral induction. {ECO:0000269|PubMed:14573523}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
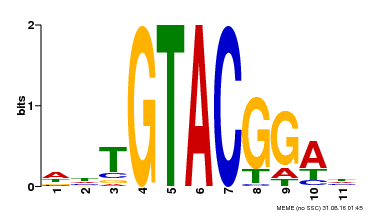
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013610197.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156 and miR157. {ECO:0000305|PubMed:12202040}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013610197.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 13A | ||||
| Refseq | XP_013678642.1 | 0.0 | squamosa promoter-binding-like protein 13A | ||||
| Swissprot | B9DI20 | 0.0 | SP13A_ARATH; Squamosa promoter-binding-like protein 13A | ||||
| Swissprot | P0DI11 | 0.0 | SP13B_ARATH; Squamosa promoter-binding-like protein 13B | ||||
| TrEMBL | A0A0D3E9W5 | 0.0 | A0A0D3E9W5_BRAOL; Uncharacterized protein | ||||
| TrEMBL | A0A3P6E835 | 0.0 | A0A3P6E835_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g097630.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4111 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 1e-167 | SBP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



