 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013599256.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 410aa MW: 45521.7 Da PI: 9.5705 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 47 | 5.8e-15 | 332 | 387 | 5 | 59 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH.HHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL.kkeleelkkevakl 59
+r++r++kNRe+A rsR+RK+a++ eLe kv++L++ N++L +k++e ++k+ ++l
XP_013599256.1 332 RRQKRMIKNRESAARSRARKQAYTLELEAKVAQLKEMNEELhRKQVEIIEKQKKQL 387
69**************************************9556677777776666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 6.5E-15 | 328 | 393 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.484 | 330 | 375 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 4.69E-10 | 332 | 376 | No hit | No description |
| Pfam | PF00170 | 3.5E-13 | 332 | 389 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 6.7E-14 | 332 | 388 | No hit | No description |
| CDD | cd14707 | 7.06E-24 | 332 | 386 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 335 | 350 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 410 aa Download sequence Send to blast |
MGSRMNFVDV VRDEVINAKQ PALGSGLPLT RQNSVFSLTF DEFQNSWGGG IGKDFGSMNM 60 DELLKNIWTA EESHSMMANN TVMNSGGGLS VGVGGEVGGC GLQRQGSIAL PRTISQKRVD 120 DVWKELLKEE DDTGASGVPQ RQQTLGEMTL EEFLLKAGVV REEPQQHVER LDNFNGGFYG 180 FGSNAGLGSA PNEFGPNQPY GVTVRPDMLK IQTQPLQMQQ RQQLIPKQVE PFPKQTTVAF 240 SNTVGLDNRS QPPTQNQEVK PSILGVRPMN NNNLQAVDFK TGVTVGEVSP GSQMSPDLTP 300 KSNMDASLSP PVPYMFGRAR KTGAVLEKVI ERRQKRMIKN RESAARSRAR KQAYTLELEA 360 KVAQLKEMNE ELHRKQVEII EKQKKQLLEP MHQPWGCKRQ CLRRTSTGPW |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots and flowers. {ECO:0000269|PubMed:11884679}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
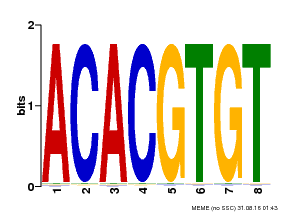
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013599256.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX122893 | 0.0 | JX122893.1 Brassica napus ABA-responsive element binding factor 3 (ABF3.1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013599256.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 isoform X1 | ||||
| Refseq | XP_013599257.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 isoform X1 | ||||
| Refseq | XP_013599258.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 isoform X1 | ||||
| Refseq | XP_013599259.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 6 isoform X1 | ||||
| Swissprot | Q9M7Q3 | 0.0 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | X2C489 | 0.0 | X2C489_BRANA; ABA-responsive element binding factor 3 | ||||
| STRING | Bo7g116570.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 1e-176 | abscisic acid responsive elements-binding factor 3 | ||||



