 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013586359.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 841aa MW: 96358.3 Da PI: 7.1942 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 79.1 | 8.2e-25 | 84 | 187 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk..............tekerrtraetrtgCkaklkvkkekdgkwevtkle 82
fY+eY++++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ +++ + +r+ +t+Cka+++vk++ dgkw++++++
XP_013586359.1 84 FYQEYSRTIGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDKSfnrprarqskqdhpENNMSGRRTCAKTDCKASMHVKRKPDGKWVIHSFV 178
********************************************999****9998887766666669999************************* PP
FAR1 83 leHnHelap 91
+HnH+l p
XP_013586359.1 179 RDHNHDLLP 187
******975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 9.1E-23 | 84 | 187 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.1E-25 | 285 | 376 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.124 | 565 | 601 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.9E-6 | 576 | 603 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 841 aa Download sequence Send to blast |
MDIDLRLHSG DLCKEGDDEE HRGLDNEDID ISKVEDDVTV EVHTDNNSTA AGITLPEHNT 60 QQQGVNLEPL NGMEFNSHGE AYTFYQEYSR TIGFNTAIQN SRRSKTTREF IDAKFACSRY 120 GTKREYDKSF NRPRARQSKQ DHPENNMSGR RTCAKTDCKA SMHVKRKPDG KWVIHSFVRD 180 HNHDLLPAQA VSEQTRKIYA AMAKQFAEYK TVISLKSESK TSFEKGRSLS VETGDFKVLL 240 DFLSRMQILN SNFFYAVDLG DDHRVKNVFW VDAKSRVNYG SFCDVVSLDT TYVRNKYKMP 300 LAIFVGVNQH YQYMVLGCAL LSDESAATFS WLMETWLRAM GGHAPKVLIT EQDPLLNSIV 360 PEVFPNTQHC LFLWHVLIKV SENLGQVAKQ HENFTAKFEK CIYKSWKEED FARRWWKILA 420 RFGLKDDQWM QSLYEDRRKW APTYMTDVLL AGMSTSLRAD SVNAFFDKYL HKKTSVQEFV 480 KLYDTILQDR FEEEAKADSE TWNKPPAMKS PSPFEKSVSD VYTPAVFKKF QIEVLGAIAC 540 SPREENRDAT CSAFRVQDFE NNQDFVVTWS QAKAEVSCMC RLFEYKGYLC RHTLNVLQCC 600 HLSSIPSQYI LKRWTKDARS QHCPGEPQQL QTRLQRYNDL CQRALKLSEE ASISQESYNI 660 AFLAIEEALG NCAGVNTSGR SLPDVVASPT QGLISVEEDN QSRSAVKTTS KKKNPTKKRK 720 VNSEQEVIPV AAPESLQQMD KLSPRTVGLE SYYGTQQSVQ NLVQLNLMAP NRDNFYGNQQ 780 TIQGLRQLNS IAPSYDSYYT AQQGIHGQGV DFFRPPNFTY DIRDDPNVRT TQLHEDASRH 840 T |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
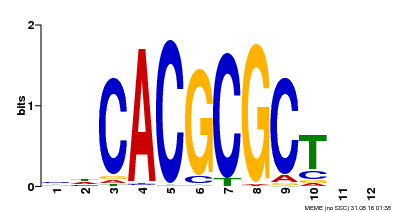
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013586359.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013586359.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A0D3CHB7 | 0.0 | A0A0D3CHB7_BRAOL; Uncharacterized protein | ||||
| STRING | Bo5g098280.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



