 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013584294.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 342aa MW: 39910.3 Da PI: 5.827 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 177.1 | 4.8e-55 | 7 | 136 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvkaeek.ewyfFskrdkkyatgkrknratksgyWkatgkdkev 92
+ppGfrFhPtdeelv++yL++kv+++++e+ ++ik++d+yk+ePwdL++ k+ +ee+ +wyfFs++dkky+tg+r+nratk g+Wkatg+dk++
XP_013584294.1 7 VPPGFRFHPTDEELVDYYLRRKVASNRIEI-DFIKDIDLYKIEPWDLQElcKIGHEEQsDWYFFSHKDKKYPTGTRTNRATKVGFWKATGRDKAI 100
69****************************.99**************9544444444359*********************************** PP
NAM 93 lskkgelvglkktLvfykgrapkgektdWvmheyrle 129
+ +++l+g++ktLvfykgrap+g+k+dW+mheyrle
XP_013584294.1 101 YL-RHSLIGMRKTLVFYKGRAPNGQKSDWIMHEYRLE 136
**.8899****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.09E-59 | 6 | 156 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.882 | 7 | 156 | IPR003441 | NAC domain |
| Pfam | PF02365 | 5.7E-29 | 8 | 135 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MNSFSLVPPG FRFHPTDEEL VDYYLRRKVA SNRIEIDFIK DIDLYKIEPW DLQELCKIGH 60 EEQSDWYFFS HKDKKYPTGT RTNRATKVGF WKATGRDKAI YLRHSLIGMR KTLVFYKGRA 120 PNGQKSDWIM HEYRLETDEN GTPQEEGWVV CRVFKKRLTA VRRMGDYDSS PSHWYGDQLS 180 FMASELETNG PRRILPNQQQ QHQYEHQQQL PYGLNASAYI LNNPNLQCKQ ELELQYNHLQ 240 SNLAHEEQLN QGNQSFSSLY MNSGNEEAMD QVTDWRVLDK FVASQLSNED AATTSASLQN 300 NAKDTSNVEC QVDEEKDQKR VSDMRDEYAA STSSSCQIDL WK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 4e-51 | 7 | 157 | 15 | 169 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Up-regulated during xylem vessel element formation. Expressed preferentially in procambial cells adjacent to root meristem. {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:25148240}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in root, shoot and hypocotyl vascular elements, columella root caps, epidermal and cortex root cells and root-hypocotyl junctions. Observed predominantly in root imature xylem vessels (PubMed:18445131). Present in root developing xylems (PubMed:16103214, PubMed:17565617). Specifically expressed in vessels in the secondary xylem of the root-hypocotyl region, and in vessels but not in interfascicular fibers in stems (PubMed:25148240). {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:17565617, ECO:0000269|PubMed:18445131, ECO:0000269|PubMed:25148240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:16103214, PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:25148240}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00136 | DAP | Transfer from AT1G12260 | Download |
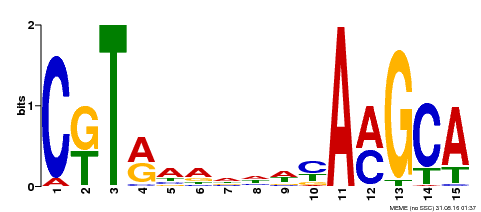
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013584294.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT029761 | 0.0 | BT029761.1 Arabidopsis thaliana At1g12260 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013584294.1 | 0.0 | PREDICTED: NAC domain-containing protein 7-like isoform X2 | ||||
| Swissprot | Q9FWX2 | 0.0 | NAC7_ARATH; NAC domain-containing protein 7 | ||||
| TrEMBL | A0A397YXX0 | 0.0 | A0A397YXX0_BRACM; Uncharacterized protein | ||||
| STRING | Bo5g017120.1 | 0.0 | (Brassica oleracea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G12260.1 | 0.0 | NAC 007 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



