 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSBRNA2T00123560001 | ||||||||
| Common Name | GSBRNA2T00123560001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 789aa MW: 86367.4 Da PI: 6.2134 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.7e-21 | 124 | 179 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
GSBRNA2T00123560001 124 KKRYHRHTPQQIQELESMFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 179
688999***********************************************999 PP
| |||||||
| 2 | START | 198.9 | 2.2e-62 | 309 | 528 | 2 | 206 |
HHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEE CS
START 2 laeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....kaet 81
la++a++el+k+a+++ep+Wvks +++n +e++++f+++k +ea+r+sg+v+ ++ lve+l+d+ +W+e+++ + +t
GSBRNA2T00123560001 309 LALTAMDELLKLAQSDEPLWVKSLdgerDELNHEEYMRTFSSTKPngLVTEASRTSGMVIINSLALVEKLMDSD-RWTEMFPcivaRSTT 397
6899************************************99999999**************************.*************** PP
EEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEE CS
START 82 levissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghsk 163
++vis g ga+qlm+aelq+lsplv R++ f+R+++q+ +g+w++vdvS+d ++++ s v+R +lpSg++++++sng+sk
GSBRNA2T00123560001 398 TDVISGGmagtrnGAIQLMNAELQVLSPLVRvRNVNFLRFCKQHAEGVWAVVDVSIDTVRENSGgSTVVIR--RLPSGCVVQDMSNGYSK 485
***************************************************************76666666..***************** PP
EEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 164 vtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
vtwveh++++++++h l+r+l++sgl +g ++wvatlqrqce+
GSBRNA2T00123560001 486 VTWVEHAEYDENQIHHLYRPLLRSGLCFGSQRWVATLQRQCEC 528
*****************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-20 | 111 | 181 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-22 | 111 | 181 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.281 | 121 | 181 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 122 | 185 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.45E-18 | 124 | 181 | No hit | No description |
| Pfam | PF00046 | 2.0E-18 | 124 | 179 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 156 | 179 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.348 | 299 | 531 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.74E-30 | 303 | 528 | No hit | No description |
| CDD | cd08875 | 3.41E-111 | 303 | 527 | No hit | No description |
| SMART | SM00234 | 1.1E-36 | 308 | 528 | IPR002913 | START domain |
| Pfam | PF01852 | 2.0E-55 | 309 | 528 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.65E-21 | 548 | 761 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 789 aa Download sequence Send to blast |
MNFGSLFDNT TGGVPTGLAY GNHTTASTVI PGGAMSHIAA ASLFSPPINT KSIYSSSGLS 60 LALEQPERGT SMRNNNGAGG DIFDGSGKNR RSREEEHESR SGSDNVEGIS GEDQDADDNK 120 PPKKKRYHRH TPQQIQELES MFKECPHPDE KQRLELSKRL CLETRQVKFW FQNRRTQMKT 180 QLERHENALL RQENDKLRAE NMSIREAMRN PTCNICGGPA MLGDVSIEEH HLRVENARLK 240 DELDRVFNLT GKFLGHHHNN HTSSLELGVG TNNNGGNFAF PPDFNGGLPP PQQATVINGV 300 DQRSVLLDLA LTAMDELLKL AQSDEPLWVK SLDGERDELN HEEYMRTFSS TKPNGLVTEA 360 SRTSGMVIIN SLALVEKLMD SDRWTEMFPC IVARSTTTDV ISGGMAGTRN GAIQLMNAEL 420 QVLSPLVRVR NVNFLRFCKQ HAEGVWAVVD VSIDTVRENS GGSTVVIRRL PSGCVVQDMS 480 NGYSKVTWVE HAEYDENQIH HLYRPLLRSG LCFGSQRWVA TLQRQCECLA ILMSSSVTSH 540 DDTSISPGGR KSMLKLAQRM TINFCSGISA PSVHSWSKLT VGNVDPDVRI MTRKSVDDPS 600 EAPGIVLSAA TSVWLPASPQ RLFDFLRNER MRCEWDILSN GGPMQEMAHI AKGQDQGNSV 660 SLLRSNPMNA NQSSMLILQE TCIDASGALV VYAPVDIPAM HVVMNGGDSS YVALLPSGFA 720 VLPDGGFDGV SGDGEQRPVG GGSLLTVAFQ ILVNNLPTAK LTVESVETVN SLISCTVQKI 780 RTALQCEN* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bna.6650 | 0.0 | seed | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
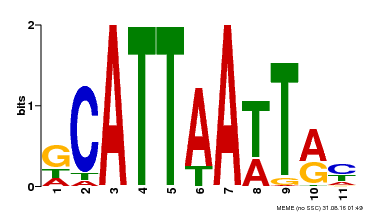
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GSBRNA2T00123560001 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013612299.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A3P6D939 | 0.0 | A0A3P6D939_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g002480.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



