 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSBRNA2T00098954001 | ||||||||
| Common Name | GSBRNA2T00098954001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 349aa MW: 39771.4 Da PI: 9.5593 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 93.7 | 8.4e-30 | 124 | 173 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krien++ rqvtf+kRrng+lKKA+ELSvLCdaeva++ifs++g+lyey+
GSBRNA2T00098954001 124 KRIENTTSRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYA 173
79***********************************************8 PP
| |||||||
| 2 | K-box | 113.9 | 1.6e-37 | 195 | 287 | 7 | 99 |
K-box 7 ksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrk 96
s++ea+++++qqe++kL+++i+ +q+++Rh++Ge+L+sL++keL++Le +Lek+++++RskKnell+++ie++qk+e elq++n++Lr+
GSBRNA2T00098954001 195 PSVTEANTQYYQQEASKLRRQIRDIQNSNRHIVGESLGSLNFKELKNLEGRLEKGISRVRSKKNELLMAEIEYMQKREMELQHDNMYLRA 284
45899************************************************************************************* PP
K-box 97 kle 99
k+
GSBRNA2T00098954001 285 KIS 287
986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 32.113 | 116 | 176 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 4.4E-38 | 116 | 175 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.51E-41 | 118 | 190 | No hit | No description |
| PRINTS | PR00404 | 1.2E-31 | 118 | 138 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 2.35E-31 | 118 | 188 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 118 | 172 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 9.7E-26 | 125 | 172 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.2E-31 | 138 | 153 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.2E-31 | 153 | 174 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.1E-27 | 200 | 286 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.643 | 202 | 292 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MIMTIRMIGH HLMDSLISCS LIKLSSHHFF SFLDPWSQIG LLKIRLTAKR GPTQEGKDKY 60 DVLGLEFSVE HVIKGCKSGF PTKNKLYGTK IEERIKIEES MEEGGSSHDA ESSKKLVRGK 120 IEIKRIENTT SRQVTFCKRR NGLLKKAYEL SVLCDAEVAL VIFSTRGRLY EYANNSVKGT 180 IERYKKACSD AVNPPSVTEA NTQYYQQEAS KLRRQIRDIQ NSNRHIVGES LGSLNFKELK 240 NLEGRLEKGI SRVRSKKNEL LMAEIEYMQK REMELQHDNM YLRAKISQGA RLNPEQQDSS 300 VIQGTAVYES GLSSHDQSQH YNRNYIPVNL LEPNQQFSGQ DQPPLQLV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1egw_A | 2e-17 | 118 | 184 | 2 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_B | 2e-17 | 118 | 184 | 2 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_C | 2e-17 | 118 | 184 | 2 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_D | 2e-17 | 118 | 184 | 2 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1tqe_P | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_Q | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_R | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_S | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_A | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_B | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_C | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_D | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_E | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_F | 2e-17 | 118 | 184 | 3 | 69 | Myocyte-specific enhancer factor 2B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Interacts genetically with TT16/AGL32 in a partially antagonistic manner during flower development. Is essential for the coordination of cell divisions in ovule, seed coat development and endosperm formation (PubMed:27776173). {ECO:0000269|PubMed:27776173}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00093 | SELEX | Transfer from AT3G58780 | Download |
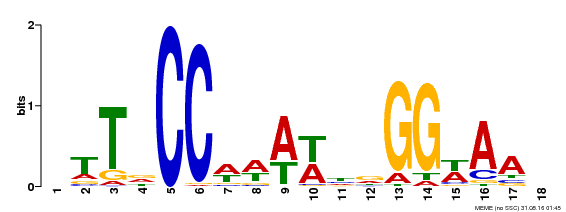
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GSBRNA2T00098954001 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JQ973085 | 0.0 | JQ973085.1 Brassica napus Shatterproof (SHP1b) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009104172.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL1 | ||||
| Swissprot | P29381 | 1e-176 | AGL1_ARATH; Agamous-like MADS-box protein AGL1 | ||||
| TrEMBL | A0A3P6BT97 | 0.0 | A0A3P6BT97_BRACM; Uncharacterized protein | ||||
| STRING | Bra003356.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3341 | 23 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G58780.1 | 1e-176 | MIKC_MADS family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



