 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSBRNA2T00082557001 | ||||||||
| Common Name | GSBRNA2T00082557001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 269aa MW: 30994.2 Da PI: 6.4604 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.7 | 4e-18 | 70 | 117 | 9 | 56 |
HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 9 keqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
eq+++Le+ Fe ++++ e++ +LAk lg++ rq+ +WFqNrRa++k
GSBRNA2T00082557001 70 LEQVKVLEKSFELGNKLDPERKIQLAKALGMQPRQIAIWFQNRRARWK 117
58*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 125.7 | 2.1e-40 | 63 | 155 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekk+rl eqvk+LE+sFe +kL+perK +la++Lg+qprq+a+WFqnrRAR+kt+qlE+dy++Lk+++d+lk++n++L +++++L +e
GSBRNA2T00082557001 63 EKKKRLPLEQVKVLEKSFELGNKLDPERKIQLAKALGMQPRQIAIWFQNRRARWKTRQLERDYDSLKKQFDSLKSDNDSLLAHNKKLLAE 152
69*************************************************************************************999 PP
HD-ZIP_I/II 91 lke 93
+++
GSBRNA2T00082557001 153 VMS 155
875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.12E-19 | 58 | 121 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 9.7E-19 | 62 | 123 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50071 | 16.082 | 63 | 119 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.99E-17 | 64 | 120 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-19 | 67 | 127 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 2.4E-15 | 70 | 117 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 2.8E-5 | 90 | 99 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 94 | 117 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.8E-5 | 99 | 115 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.9E-16 | 120 | 159 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 269 aa Download sequence Send to blast |
FQQLHEDISP DQLPSCLPPH PFNGGGNYMM NRSMSLTNVR DDPHQSVDEE NLSDDGSHMM 60 LGEKKKRLPL EQVKVLEKSF ELGNKLDPER KIQLAKALGM QPRQIAIWFQ NRRARWKTRQ 120 LERDYDSLKK QFDSLKSDND SLLAHNKKLL AEVMSLKNKD YNEASIIKRE AEASWSNNGS 180 TENSSDINLE IPRETTTTHV NTIKNLFPSS IPSSTHHQDH HGDHHHQNNE MVQEESFCNM 240 FNGIDETTSA GYWGWSDPNY NHHHHQFN* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed 3 day after germination (DAG) in first rosette leaf primordia around the veins in the apical part of the leaf and close to the midvein in the basal part of the leaf. At later stage, expressed in new forming veins, and veins and fascicular cambium of mature leaves. {ECO:0000269|PubMed:12644682}. | |||||
| Uniprot | TISSUE SPECIFICITY: Widely expressed. {ECO:0000269|PubMed:16055682}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
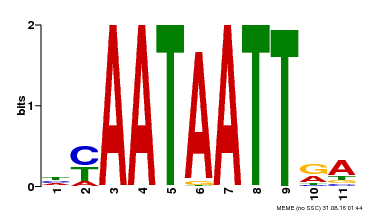
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GSBRNA2T00082557001 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013584528.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-20-like | ||||
| Swissprot | Q8LAT0 | 1e-157 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A078JQF3 | 0.0 | A0A078JQF3_BRANA; BnaCnng61510D protein (Fragment) | ||||
| STRING | Bo5g156340.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3095 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 1e-154 | homeobox protein 20 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



