 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi5g26110.2.p | ||||||||
| Common Name | BRADI_5g26110 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 330aa MW: 34067.8 Da PI: 6.1817 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 22.5 | 2.7e-07 | 22 | 70 | 2 | 45 |
SSS-HHHHHHHHHHHHHTTTT......-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGgg......tWktIartmgkgRtlkqcksrwq 45
g WT e ++ + +av+ G + W+ a+ + g+t+++++ ++
Bradi5g26110.2.p 22 GGWTREQEKAFENAVAAAGEEapegdaAWEEMAAAVE-GKTAEEVRRHYE 70
67***********************************.**********95 PP
| |||||||
| 2 | Myb_DNA-binding | 21.9 | 4.1e-07 | 170 | 205 | 12 | 47 |
HHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 12 lvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++ + +++G+g+W++I+r + Rt+ q+ s+ qky
Bradi5g26110.2.p 170 FLLGLEKYGKGDWRSISRNFVISRTPTQVASHAQKY 205
777899***************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 4.9E-4 | 20 | 75 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 6.679 | 22 | 73 | IPR017877 | Myb-like domain |
| SuperFamily | SSF46689 | 1.15E-8 | 23 | 79 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.96E-5 | 24 | 73 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 9.9E-6 | 129 | 206 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.30E-5 | 137 | 206 | No hit | No description |
| SMART | SM00717 | 0.066 | 164 | 208 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 9.3E-11 | 169 | 208 | IPR006447 | Myb domain, plants |
| SuperFamily | SSF46689 | 1.29E-9 | 169 | 211 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.3E-5 | 170 | 205 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 4.739 | 174 | 206 | IPR017877 | Myb-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 330 aa Download sequence Send to blast |
MAVEEASSSG GEEGSGSGSG SGGWTREQEK AFENAVAAAG EEAPEGDAAW EEMAAAVEGK 60 TAEEVRRHYE LLVEDVDGIE SGRVPLLVYA GDAAAAEEGG SGGGGGGKKG GGGGGGGHGE 120 KVSAAKSAEQ ERRKGIAWTE DEHRRCDQGT SIAVFFIKRK VDSNLGAWLF LLGLEKYGKG 180 DWRSISRNFV ISRTPTQVAS HAQKYFIRLN SMNRERRRSS IHDITSVNNG DTSAAQGPIT 240 GTNGQVANPG KSPKQPLQPA NPPPGVDAYG TTIGQPVGGP LVSAVGTPVS LPVPAPPHLA 300 YGMHAPVPGT VVPRAPINMP YPMPPPSSR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 7e-14 | 16 | 89 | 2 | 73 | RADIALIS |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 105 | 113 | GGKKGGGGG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00500 | DAP | Transfer from AT5G08520 | Download |
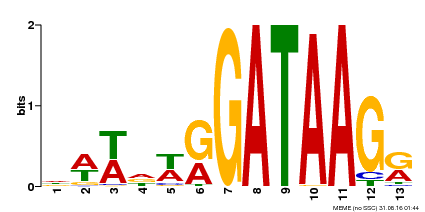
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi5g26110.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP093944 | 1e-135 | FP093944.1 Phyllostachys edulis cDNA clone: bphylf060p06, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024311422.1 | 0.0 | transcription factor SRM1 isoform X1 | ||||
| Swissprot | Q9FNN6 | 5e-90 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A0Q3IGB1 | 0.0 | A0A0Q3IGB1_BRADI; Uncharacterized protein | ||||
| STRING | BRADI5G26110.1 | 1e-162 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6376 | 38 | 54 | Representative plant | OGRP275 | 17 | 122 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G08520.1 | 9e-71 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi5g26110.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



