 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi2g59480.1.p | ||||||||
| Common Name | BRADI_2g59480, LOC100834181 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 815aa MW: 90695.5 Da PI: 6.2665 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.1 | 6.8e-23 | 127 | 228 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldg 86
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d++g W++++i+r++++r++l++GW+ Fv++++L +gD ++F +
Bradi2g59480.1.p 127 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHGMDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--R 215
99*****************************7555544..58************************************************..4 PP
SSEE..EEEEE-S CS
B3 87 rsefelvvkvfrk 99
+++el+v+v+r+
Bradi2g59480.1.p 216 GESGELRVGVRRA 228
49999*****996 PP
| |||||||
| 2 | Auxin_resp | 109.2 | 4.2e-36 | 253 | 334 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++s+G Rf+m+fe+e+++e+r++Gt++g ++ldp Wp+S WrsLk
Bradi2g59480.1.p 253 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSIGVRFRMRFEGEEAPEQRFTGTIIGSENLDP-LWPESSWRSLK 334
89*********************************************************************.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.14E-40 | 118 | 256 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.5E-39 | 121 | 242 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.08E-19 | 125 | 227 | No hit | No description |
| PROSITE profile | PS50863 | 11.702 | 127 | 229 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 9.5E-22 | 127 | 229 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.8E-21 | 127 | 228 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.4E-33 | 253 | 334 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 4.0E-12 | 675 | 778 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 1.44E-11 | 687 | 770 | No hit | No description |
| PROSITE profile | PS51745 | 23.755 | 693 | 786 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 815 aa Download sequence Send to blast |
MPPVAMAPPP QGPSAGDPLF NELWHACAGP LVTVPRVGDL VFYFPQGHIE QVEASMNQVA 60 DNQMRLYDLP SKLLCSVINV ELKAEADTDE VYAQVMLIPE NDQNEMAVEK SSSKAATTLA 120 KPAVRSFCKT LTASDTSTHG GFSVLRRHAD ECLPPLDMTQ SPPTQELVAK DLHGMDWRFR 180 HIFRGQPRRH LLQSGWSVFV SSKRLVAGDA FIFLRGESGE LRVGVRRAMR QLSNVPSSVI 240 SSHSMHLGVL ATAWHAINTK SMFTVYYKPR TSPSEFIIPY DQYMESVKNN YSIGVRFRMR 300 FEGEEAPEQR FTGTIIGSEN LDPLWPESSW RSLKVRWDEP STIPRPDRVS PWKIEPASSP 360 PVNPLPLSRV KRPRPNVPPA SPESSALTKE GATKVDVDSA QAQRNQTSMV LQGQEPMTLR 420 SNNLTDSNDS DATVQKPMMW SPSPNIGKSR PLTFQQRPSM DNWMQLGRRE TDYKDASSGA 480 QSFGDSPGFF VQTYDEAPNH LASFKNQFQD QGPARHFSEP YFFMHQQPNL TVDSSTKMHP 540 ETNELHFWNG QNTVYGHSGD QSQGFRFEEH PSNWSSQQFS RVEQPRVIRP HASIAPVDLE 600 KTREGSGFKI FGFKVDTASA PTNHLSSPMA ATHEPALQTQ PSASLNQLQH AQTDCFPEVS 660 VSTGGTNENE KSIQQAPQSS KDVQSKSHGA STRSCTKVHK QGVALGRSVD LSKFSDYDEL 720 KAELDKMFEF DGELMSSNKN WQIVYTDNED DMMLVGDDPW GEFCSIVRKI CIYTKEEVQK 780 MNSKLSAPRK EDGAAPCDGA NENAALPAPG NLDN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-167 | 12 | 359 | 12 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-167 | 12 | 359 | 12 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-167 | 12 | 359 | 12 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-167 | 12 | 359 | 12 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-167 | 12 | 359 | 12 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 370 | 374 | KRPRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
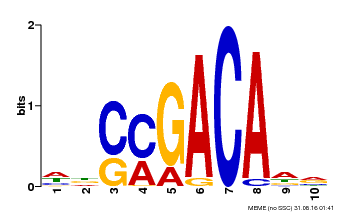
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi2g59480.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003564986.1 | 0.0 | auxin response factor 4 | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A0Q3JFT4 | 0.0 | A0A0Q3JFT4_BRADI; Auxin response factor | ||||
| STRING | BRADI2G59480.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 | Representative plant | OGRP550 | 15 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi2g59480.1.p |
| Entrez Gene | 100834181 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



