 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi2g58130.4.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1204aa MW: 132739 Da PI: 8.3086 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.4 | 0.001 | 1159 | 1185 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ C+++F+ s++ rH r+ H
Bradi2g58130.4.p 1159 YVCHepGCTQTFRFVSDFSRHKRRtgH 1185
99********************98666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 3.8E-14 | 28 | 69 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.827 | 29 | 70 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 4.7E-13 | 30 | 63 | IPR003349 | JmjN domain |
| SMART | SM00558 | 5.0E-47 | 196 | 365 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 36.963 | 199 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 6.32E-27 | 211 | 363 | No hit | No description |
| Pfam | PF02373 | 3.8E-37 | 229 | 348 | IPR003347 | JmjC domain |
| SMART | SM00355 | 31 | 1076 | 1098 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.073 | 1099 | 1123 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 1099 | 1128 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.9E-5 | 1100 | 1127 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1101 | 1123 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.05E-9 | 1115 | 1155 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-8 | 1128 | 1152 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.016 | 1129 | 1153 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.365 | 1129 | 1158 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1131 | 1153 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.08E-8 | 1147 | 1182 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-8 | 1153 | 1181 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.3 | 1159 | 1185 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.221 | 1159 | 1190 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1161 | 1185 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1204 aa Download sequence Send to blast |
MMSSSSSPPP PARVAAELVP PWLKSLPLAP EFHPTAAEFA DPVAYILKIE PAAEPFGICK 60 VVPPCPQPPK KTTLSNLTRS FAALHPDDPA PTFPARHQQL GLSPRSRRPA LTAVWQSSRS 120 YTLPQFESRA AASRKTLLAR LNVPASKHLS PLEHEALFWS ASADRPVTVD YASDMPGSGF 180 SAPSTCAARP PSQQAHVGET AWNMRGAARS PGSLLRFMRD EVPGVNTPML YVGMTFSWFA 240 WHVEDHDLHS LNYMHFGAAK TWYGVPRDAA LAFEDVVREH GYGGEVNPLE TFATLGKKTT 300 VMSPEVLVGL GVPCCRLVQN EGDFVVTFPG SYHCGFSHGF NCGEASNIAT PEWLRVAKEA 360 AIRRASINRP PMLSHYQLLY ELALSMCIRD PSIGPMEPRS SRLKEKKKGE GGQLVKKIFV 420 QNAIEDNELL SSLLNDGSSC IILPINADDG PVLSALRSRS QLKAKSNTSD GLCSSGEALE 480 ESRCLSETFD RNGEIINCGA LSSSKESPSS VCSGSMHDCV NLSCSSDTHN AEGDKVDVIS 540 AAGLLDQGLL SCVSCGILSF SCVAVIKPRE CTSKYLMSSD YNLINDQLVN SGRIHPANAT 600 SEGTDGGILH SDEDIQNKKI KVCHDCSELS RHMTESQHND SSHVCFDGTK MSSSSIKCQE 660 RPSSQSSQCI GGSGILNGPR GFRTRNKYLL KMALSEGFQP NNIYQSMEKK GQSEPSNSKK 720 IVKEPLVTGG TDYDARCKST AIATGDPRSS TATINISDQP IVEFDKDSSR MHVFCLEHAV 780 EVEKRLQAIG GAHIILLCRP EYLKIEAEAR TLAAEMKVEY DWKDIHFREA NMEDRDMIQE 840 VLQDEETIPT NSDWAVKLVN LKTCERNQGR QKKVVFAGRW CGRVWMSNQV HPYLARRIES 900 HELEEIDNSS GVEVSKRIGS TITAVMRSSK KRENMILKET NSKRIKQTQE CNSGALKGVA 960 EVPSPSPAGV VLRVSSRIAN RANKIKSEMT EEEDPAGRPK SEVTCPSRPG PSKQKTKVKA 1020 KNQIMPPTAS LKDEKEHPSA TKGSSVSCYT KQRTRTEAKE ATGETGTPRD PKQEEYVCNI 1080 DGCLMSFDTK KELSLHKHNI CPVKGCGKKF FVHRYLLQHR KVHTDDRPLK CPWEGCDVAF 1140 KWTWARTEHL RVHTGDRPYV CHEPGCTQTF RFVSDFSRHK RRTGHSTKKT NKHKSNQGSS 1200 KLW* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 5e-72 | 22 | 386 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-72 | 22 | 386 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
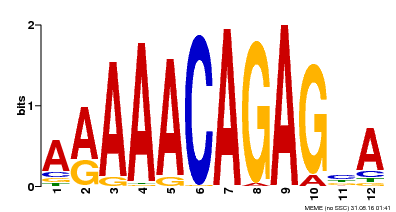
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi2g58130.4.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010232711.2 | 0.0 | lysine-specific demethylase JMJ705 | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | A0A2K2DGL8 | 0.0 | A0A2K2DGL8_BRADI; Uncharacterized protein | ||||
| STRING | BRADI2G58130.1 | 0.0 | (Brachypodium distachyon) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-174 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi2g58130.4.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



