 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi2g04810.1.p | ||||||||
| Common Name | BRADI_2g04810, LOC100822315 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 495aa MW: 52164.7 Da PI: 6.0492 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.3 | 1.3e-31 | 278 | 333 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++ LGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Bradi2g04810.1.p 278 KARRCWSPELHRRFVNALQILGGAQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 333
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.652 | 275 | 335 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-28 | 276 | 336 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.58E-17 | 276 | 336 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.9E-26 | 278 | 333 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-6 | 280 | 331 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 495 aa Download sequence Send to blast |
MDSSPSDLTL DYTPNGNGNG GGGGGHPMTP KQAPLVVEQH HLTAAEQASA TTTVQKLQEF 60 LSRLDDERLK IDAFKRELPL CMQLLNQAME AYRQQLEACQ MGSSHGGTAG PGRAPLVLEE 120 FIPLSKNIDA ADQNQQAASE NPKASWMVSA QLWNGPPPSS ADAPQTPKER SEHPPLDAGT 180 RGNGGAFLPF TKEMITTTAD HSPAALLPEL ALAPGAEKDA LVGINNGLSA SGDKKPYLHH 240 DAGGGGNNGL AVARRDDDVL QNGSQTTTAA AAPQTNRKAR RCWSPELHRR FVNALQILGG 300 AQVATPKQIR ELMKVDGLTN DEVKSHLQKY RLHTRRPMQA PAAPPAGAPQ LVVLGGIWMP 360 PDYAAQAAQA AAGPAAIYGA HPATQTHYTA AVSSAAAQEY YHSAAAHHHH LQQQQHHHPH 420 PAMAHGRAMA PPAAAYKVHH PVAASPESEG RGSGGGGGGG RERSESIEEE GEGEDDEDDE 480 DAIAADAEAE EIKY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 2e-13 | 278 | 331 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-13 | 278 | 331 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-13 | 278 | 331 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-13 | 278 | 331 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may play a role in regulatory networks controlling development and metabolism. {ECO:0000250|UniProtKB:Q6Z869}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
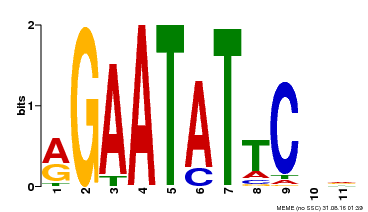
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi2g04810.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT086572 | 1e-100 | BT086572.2 Zea mays full-length cDNA clone ZM_BFc0184N23 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003565416.2 | 0.0 | transcription factor NIGTH1 | ||||
| Swissprot | Q5VRW2 | 1e-120 | NOH1_ORYSJ; Transcription factor NIGTH1 | ||||
| TrEMBL | I1HCK2 | 0.0 | I1HCK2_BRADI; Uncharacterized protein | ||||
| STRING | BRADI2G04810.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9773 | 33 | 38 | Representative plant | OGRP4303 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 2e-50 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi2g04810.1.p |
| Entrez Gene | 100822315 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



