 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi1g71810.2.p | ||||||||
| Common Name | BRADI_1g71810, LOC100845729 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1035aa MW: 115552 Da PI: 6.0409 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176 | 4.9e-55 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenpt 93
lke ++rwl++ ei++iL+n+ +++++ e+++rp sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een +
Bradi1g71810.2.p 19 LKEaQHRWLRPAEICEILKNYGNFRIAPEPPNRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKSGSIDVLHCYYAHGEENIN 111
45559**************************************************************************************** PP
CG-1 94 fqrrcywlLeeelekivlvhylevk 118
fqrr+yw+Lee++ +ivlvhylevk
Bradi1g71810.2.p 112 FQRRSYWMLEEDFMHIVLVHYLEVK 136
**********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.389 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.3E-76 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.6E-48 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| CDD | cd00102 | 0.00802 | 445 | 532 | No hit | No description |
| Gene3D | G3DSA:2.60.40.10 | 3.6E-5 | 445 | 532 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 2.5E-5 | 446 | 530 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 3.97E-16 | 446 | 531 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 5.13E-17 | 636 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 7.0E-18 | 640 | 746 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.52E-13 | 644 | 738 | No hit | No description |
| PROSITE profile | PS50297 | 17.794 | 646 | 740 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.79 | 646 | 678 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 2.1E-6 | 651 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.033 | 679 | 708 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.63 | 679 | 711 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 330 | 718 | 747 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 5.31E-8 | 850 | 903 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.12 | 852 | 874 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.663 | 853 | 882 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0034 | 854 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0013 | 875 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.487 | 876 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0017 | 877 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0001077 | Molecular Function | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1035 aa Download sequence Send to blast |
MAEGRRYAIA PQLDIEQILK EAQHRWLRPA EICEILKNYG NFRIAPEPPN RPASGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKSGSIDVLH CYYAHGEENI NFQRRSYWML 120 EEDFMHIVLV HYLEVKAGKS SSRTREHDNM LQGARVDSPL SQLPSQTTDG ESSLSGQASE 180 YEETESDIYS GGAGYHSISG MQQHENGAGP IIDASFYSSY VPASSVGNHQ GLQATATNTG 240 FYSYDQDNLP VVPNESGHGI PFNGPNGQFD LSSWNEMTKP DKGIHQMPPY GTHVPPEQSP 300 FTEVPGIESF TFDEVYSNGL GIKDNSHADT DAEPLWQLPS AIGGSFATVD SFQQINGFLE 360 EAINYPLLKT QSSNLSDILK DSFKKSDSFT RWMTKELADV DDSQIKPSSE YWNSEDADNI 420 IGASSHDQLD QFTLGPMLAQ DQLFSIIDFS PSWAYAGAKT RILVTGKFLK PDEVIRFKWS 480 CMFGEIEVPA EILADGTLGC YSPSQKTGRV PFYVTCSNRL ACSEVREFEY RPSNSQYMDA 540 PSLHGARNKT YLQMRLDKLL SLGPDEFHAT LSNNTKELID LNRKINLLMK NNDSWSELLK 600 LAGDNELVIE DKQDQFLENC IRDKLHIWLL HKAGDGGKGP GVLDKEGQGV LHLAAALGYD 660 WAIRPTITAG VNINFRDARG WTALHWAAFC GRERTVVALI ALGAAPGALT DPSPDFPSGS 720 TPADLASSNG HKGISGYLAE SSLTCHLQTL NLKEAMGSNA SEISGLPGIG DVSERSVSPL 780 AREGLQTGSM GDSLGAVRNA AQAAARIYQV FRVQSFQRKQ AVQYEDDSGV ISDERALSLL 840 SYKTSKPGQF DPKHAAATRI QNKFRGWKGR KEFLLLRRRV VQIQAHVRGH QVRKHYRKII 900 WSVGIVEKVI LRWRRRGAGL RGFRSTEGAP DSTSSSAVDV IPNKPGEDDY SFLQEGRKQT 960 EERLQRALAR VKSMVQYPDA RDQYQRILTV VTKMQESQPM QENMLEESTE MDEGFLMSEF 1020 QELWDDDTPM PGYF* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
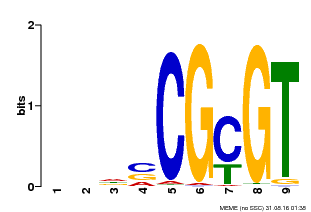
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi1g71810.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003558617.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| TrEMBL | I1H8R1 | 0.0 | I1H8R1_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G71810.2 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 | Representative plant | OGRP562 | 15 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi1g71810.2.p |
| Entrez Gene | 100845729 |



