 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi1g64687.1.p | ||||||||
| Common Name | BRADI_1g64687, LOC100838726 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 348aa MW: 37837.7 Da PI: 4.7271 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.6 | 6.1e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWT+eEd++l+ +++q+G g+W++ ++ g+ R++k+c++rw +yl
Bradi1g64687.1.p 14 RGRWTAEEDDILASYIAQHGEGSWRSMPKNAGLLRCGKSCRLRWINYL 61
8*******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.9 | 4e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ + eEd+l+v++++ lG++ W++Ia++++ gRt++++k++w+++l
Bradi1g64687.1.p 67 RGSISREEDDLIVKLHATLGNR-WSLIASHLP-GRTDNEIKNYWNSHL 112
678899****************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-22 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.042 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.1E-28 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.5E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.51E-11 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.825 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-25 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.6E-17 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.8E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.46E-13 | 71 | 112 | No hit | No description |
| Pfam | PF06640 | 1.5E-31 | 130 | 247 | IPR010588 | Myb-related protein P, C-terminal |
| Pfam | PF06640 | 1.1E-22 | 250 | 347 | IPR010588 | Myb-related protein P, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009813 | Biological Process | flavonoid biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 348 aa Download sequence Send to blast |
MGRAPCCEKV GLKRGRWTAE EDDILASYIA QHGEGSWRSM PKNAGLLRCG KSCRLRWINY 60 LRDGVKRGSI SREEDDLIVK LHATLGNRWS LIASHLPGRT DNEIKNYWNS HLSRQIHTFR 120 RTYTAGNDTT ITIDISKLSA AGKRRGGRTA GQSPKSSTKK HPEPETEPNK AKDASSPVTA 180 TSSVSSALQS DEARSAVVDP DQNQPNNSIS GSHTSDGPCS QDETGPFFMD PIDQAGGLLE 240 ANSAMDQFGL WEADSSFNQI GHLEESQMEA LLSNGVAMES GLTGVEPEGQ VQVDDLLDMD 300 WEGFAAHLWN EPAQSDMIQA AEPKTTTTGS DPDELDSFVS WLLSDAC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-23 | 12 | 118 | 5 | 110 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor postulated to regulate the biosynthetic pathway of a flavonoid-derived pigment in certain floral tissues. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
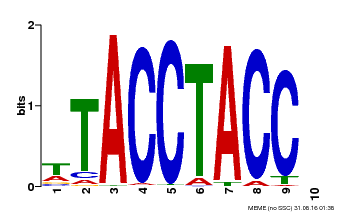
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi1g64687.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK356334 | 1e-156 | AK356334.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1033F20. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003558096.1 | 0.0 | myb-related protein P | ||||
| Swissprot | P27898 | 8e-95 | MYBP_MAIZE; Myb-related protein P | ||||
| TrEMBL | I1H6A7 | 0.0 | I1H6A7_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G64687.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47460.1 | 9e-71 | myb domain protein 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi1g64687.1.p |
| Entrez Gene | 100838726 |



