 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi1g03020.1.p | ||||||||
| Common Name | BRADI_1g03020, LOC100837762 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 147aa MW: 16068.5 Da PI: 10.5874 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 57.9 | 1.4e-18 | 26 | 60 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C+ C+ttkTplWR gp g+k+LCnaCG++yrkk++
Bradi1g03020.1.p 26 CTDCNTTKTPLWRGGPCGPKSLCNACGIRYRKKRR 60
********************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57716 | 1.43E-13 | 19 | 59 | No hit | No description |
| PROSITE profile | PS50114 | 12.848 | 20 | 56 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 2.9E-14 | 20 | 72 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 7.4E-15 | 24 | 60 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 7.90E-14 | 25 | 60 | No hit | No description |
| Pfam | PF00320 | 2.0E-16 | 26 | 60 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 26 | 51 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 147 aa Download sequence Send to blast |
MDSPVEKGSG SLDPDDRTAS GEPKACTDCN TTKTPLWRGG PCGPKSLCNA CGIRYRKKRR 60 EALGLDGPKR RETAACAHTA GEGAEQPPKK KTKREREEVT VELRMVGFGK AAVLKQRRRM 120 RRRRRLGEEE KAAILLMALS SGVIYA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 115 | 121 | QRRRMRR |
| 2 | 117 | 122 | RRMRRR |
| 3 | 117 | 124 | RRMRRRRR |
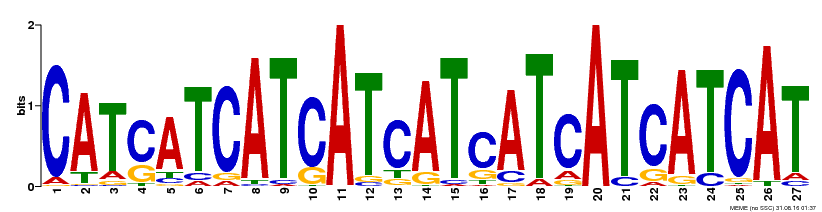
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00554 | DAP | Transfer from AT5G49300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi1g03020.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP092837 | 9e-88 | FP092837.1 Phyllostachys edulis cDNA clone: bphyst009f16, full insert sequence. | |||
| GenBank | FP101260 | 9e-88 | FP101260.1 Phyllostachys edulis cDNA clone: bphylf037m05, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003563322.1 | 1e-101 | GATA transcription factor 16 | ||||
| TrEMBL | I1GLB3 | 1e-100 | I1GLB3_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G03020.1 | 1e-101 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2418 | 38 | 92 | Representative plant | OGRP68 | 17 | 287 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G49300.1 | 1e-25 | GATA transcription factor 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi1g03020.1.p |
| Entrez Gene | 100837762 |



