 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bradi1g02760.1.p | ||||||||
| Common Name | BRADI_1g02760, LOC100835193 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Brachypodieae; Brachypodium
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 963aa MW: 105093 Da PI: 5.3586 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 127.6 | 4.7e-40 | 155 | 232 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adls ak+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Bradi1g02760.1.p 155 ACQVEGCCADLSAAKDYHRRHKVCEMHAKANTAVVGNTVQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNRRRRKTRP 232
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.6E-33 | 148 | 217 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.308 | 153 | 230 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-37 | 154 | 234 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.0E-29 | 156 | 229 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 2.18E-8 | 723 | 847 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.4E-7 | 744 | 849 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.25E-8 | 746 | 846 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 963 aa Download sequence Send to blast |
MEAAGVGSQS RRLYGGGLGE PAQDMRGKRL FGWDLNDWSW DSERFVATPA PAAEKANGLS 60 LNSSPSSSEE ADVEVARSGN VRGDSDKRKR VVVIDDDGDD QKDEDPVDNN GRVLSLRIGG 120 DTTVAGGAVE GGAVNEEDRN GKKIRVQGGS SSGPACQVEG CCADLSAAKD YHRRHKVCEM 180 HAKANTAVVG NTVQRFCQQC SRFHLLQEFD EGKRSCRRRL AGHNRRRRKT RPEIAVGGTP 240 IEDKVGSYLV LSLLGICANL NSENAEHLQG QELLSNLWRN LGTVAKSLDP KELCKLLETC 300 QSMQNGSNTG TSEAANALVN SAAVEAAGPS NSKAPFTNGG QREQTSSAVI PLQSNATVVA 360 TPETPACRIR NFDLNDTCND MEGFEDGSNC PSVQQDSTQS PPQTSGNSDS TSAQSLSSSN 420 GDAQCRTDKI VFKLFDKVPS DLPPILRSQI LGWLSSSPTD IESYIRPGCI ILTVYLRLVD 480 SAWRELSENM SLYLDKLLSS STDNFWASSL VFVMVRHQIV FMHNGQVMLD RPLAPNSHHY 540 CKVLCVSPVA APSSATVNFR VEGFNLVSAS SRLICSFEGR CIFQEDTAIV DDAAEHEDIE 600 CLNICCSLPG SRGRGFIEVE DSGFSNGFFP FIVAEQDVCS EVCELESIFK SSSHEQADND 660 NARSQALEFL NELGWLLHRA NIISKHDKVE LPLAAFNLLR FRNLGIFAME REWCAVTKVL 720 LDLLFDGFVD VGLQSPKEVV LSENLLHTAV RGKSVRMVRF LLRYKPSKDQ KEIAESYLFR 780 PDARGPSTFT PLHIAAATSD AEDVLDALTS DPGLVGLNAW KNARDETGFT PEDYARQRGN 840 DAYMDLVQKK IDKNLGEGHV VLGVPSSMCP VLTDGAKPGD ISLEICKSMP MAPQPVSRCN 900 ICSRQARMYP SSFANTFLYR PAMFTVMGVA VICVCVGILL HTLPKVYAAP NFRWELLERG 960 PM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-30 | 149 | 229 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
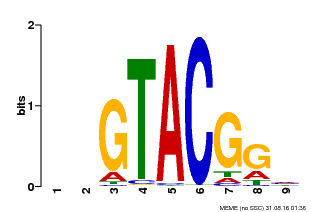
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Bradi1g02760.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF447880 | 0.0 | KF447880.1 Triticum aestivum squamosa promoter-binding-like protein 6 (SPL6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003563064.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | I1GL80 | 0.0 | I1GL80_BRADI; Uncharacterized protein | ||||
| STRING | BRADI1G02760.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 | Representative plant | OGRP2595 | 14 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Bradi1g02760.1.p |
| Entrez Gene | 100835193 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



