 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00049.51 | ||||||||
| Common Name | AMTR_s00049p00081480, LOC18441867 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 599aa MW: 67923.8 Da PI: 5.0259 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 83.3 | 2.7e-26 | 194 | 247 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++Wt +LHe+Fv+av+q+ G++k Pk+il+lm+v+gLt+e+v+SHLQkYRl
evm_27.model.AmTr_v1.0_scaffold00049.51 194 KPRVNWTVDLHEQFVDAVHQI-GIDKVGPKKILDLMHVPGLTRENVASHLQKYRL 247
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 75.4 | 2.2e-25 | 16 | 124 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeiree 68
vl+vdD+ + +++l+++l+k y +v+++ +e+al++l+e++ +D+++ D++mp+mdG++ll+
evm_27.model.AmTr_v1.0_scaffold00049.51 16 VLVVDDDLTWLKILEKMLKKCLY-KVTTCNLAEDALKMLRERKdgFDIVISDVNMPDMDGFKLLELVGL- 83
89*********************.***************999999*******************87754. PP
TTTSEEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 69 epklpiivvtahgeeedalealkaGakdflsKpfdpeelvk 109
e++lp+++++ +e++ + + +k Ga d+l Kp+ ++el++
evm_27.model.AmTr_v1.0_scaffold00049.51 84 EMDLPVLMMSVDDEKSRVMKGVKHGACDYLLKPVRMKELRN 124
558***********************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.4E-143 | 1 | 584 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 1.99E-33 | 12 | 140 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 5.8E-42 | 13 | 145 | No hit | No description |
| SMART | SM00448 | 1.6E-29 | 14 | 126 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.743 | 15 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 2.9E-22 | 16 | 125 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.74E-28 | 17 | 129 | No hit | No description |
| PROSITE profile | PS51294 | 9.958 | 191 | 250 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.78E-18 | 192 | 251 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-28 | 193 | 252 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-22 | 194 | 247 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.9E-7 | 196 | 246 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 599 aa Download sequence Send to blast |
MESVLPSSQF PNGLRVLVVD DDLTWLKILE KMLKKCLYKV TTCNLAEDAL KMLRERKDGF 60 DIVISDVNMP DMDGFKLLEL VGLEMDLPVL MMSVDDEKSR VMKGVKHGAC DYLLKPVRMK 120 ELRNIWQHVY RKKLLETKDM DFNEDFQRLR KGSDDSDEGS LFYGGDLKLS KKRKDAKDED 180 GDDKDTNDPA SVKKPRVNWT VDLHEQFVDA VHQIGIDKVG PKKILDLMHV PGLTRENVAS 240 HLQKYRLYLA RLQRQPEGTM PPNSNSREPP ESQGSNSTFS GSSDETILPQ NALSEIHDEF 300 TSLHKTVTKR ALALSKLDLL DVQKQDPVCH EGLQFISFGN TVYCEDYTSS TFPWQQHANE 360 ELPTLQIKQQ HANEKIPAMH INQQSCVGDC TNLEVPFQQP DIQIQLNHPL SSGHPYSSDF 420 IERTRMTNKG TFDLLNQTKQ PITDCKLDHV SPLGSSADSM ITFCLERNYP STKTYELESY 480 WDFPHININL QGGSTVASSM ASQFEELHLR SVQSHGHVKD FVPNGMEMIH LNDSISAAEI 540 PSFFYEETSI SSDDLFNLMD DSLLTEGMSN LSHQLISSSI SSPMLRDTWS TPINWVEM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5lxu_A | 1e-17 | 194 | 249 | 1 | 56 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
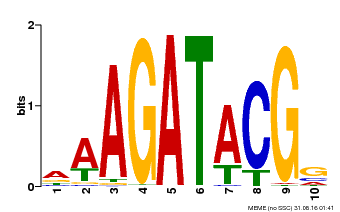
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006852154.1 | 0.0 | two-component response regulator ORR21 isoform X1 | ||||
| Refseq | XP_020527763.1 | 0.0 | two-component response regulator ORR21 isoform X1 | ||||
| TrEMBL | W1PZU7 | 0.0 | W1PZU7_AMBTC; Two-component response regulator | ||||
| STRING | ERN13621 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP292 | 17 | 120 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 9e-95 | response regulator 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00049.51 |
| Entrez Gene | 18441867 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



