 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00040.303 | ||||||||
| Common Name | AMTR_s00040p00235500, LOC18441501 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 247aa MW: 27729 Da PI: 6.1017 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 64.9 | 1.1e-20 | 51 | 103 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
++++++eq+e Le++F+++ +++ e++ +LA +lgL rqV vWFqNrRa++k
evm_27.model.AmTr_v1.0_scaffold00040.303 51 KRRLSEEQIESLEKCFQEETKLQPERKMRLAGELGLHPRQVAVWFQNRRARWK 103
4579************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 118.8 | 3e-38 | 50 | 141 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkray 70
+krrls+eq+++LE++F+ee+kL+perK++la eLgl+prqvavWFqnrRAR+k+kq+E+ y++Lk+++
evm_27.model.AmTr_v1.0_scaffold00040.303 50 RKRRLSEEQIESLEKCFQEETKLQPERKMRLAGELGLHPRQVAVWFQNRRARWKSKQVEHLYHSLKHQF 118
8******************************************************************** PP
HD-ZIP_I/II 71 dalkeenerLekeveeLreelke 93
+++e+++L++ev +Lr+el+e
evm_27.model.AmTr_v1.0_scaffold00040.303 119 HLVSKEKQKLHEEVLKLRAELRE 141
*******************9975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.68E-19 | 40 | 106 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.957 | 45 | 105 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-18 | 48 | 109 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.79E-18 | 50 | 106 | No hit | No description |
| Pfam | PF00046 | 4.9E-18 | 51 | 103 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-20 | 52 | 112 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 3.5E-5 | 76 | 85 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 80 | 103 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 3.5E-5 | 85 | 101 | IPR000047 | Helix-turn-helix motif |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010434 | Biological Process | bract formation | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 247 aa Download sequence Send to blast |
MMMPWNGNLL LPPYGGGGWS HSPSMAAVYN DGPGGGAMGA FLKQEGGTGR KRRLSEEQIE 60 SLEKCFQEET KLQPERKMRL AGELGLHPRQ VAVWFQNRRA RWKSKQVEHL YHSLKHQFHL 120 VSKEKQKLHE EVLKLRAELR ERERERGMTA REESGVVELS TVGGEESASA CVGNATTTNY 180 EDNGDEDDDT LMKCDCHPFS TINCRYGAGC SDDNNYLIPC LDDDGRPSQP NYYYNWDVGP 240 GCIKSL* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 97 | 105 | RRARWKSKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Putative transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00486 | DAP | Transfer from AT5G03790 | Download |
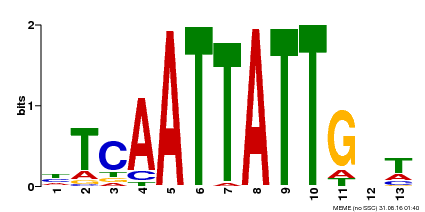
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006851793.1 | 0.0 | putative homeobox-leucine zipper protein ATHB-51 | ||||
| Swissprot | Q9LZR0 | 5e-35 | ATB51_ARATH; Putative homeobox-leucine zipper protein ATHB-51 | ||||
| TrEMBL | W1Q052 | 0.0 | W1Q052_AMBTC; Uncharacterized protein | ||||
| STRING | ERN13260 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP129 | 16 | 189 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G03790.1 | 1e-31 | homeobox 51 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00040.303 |
| Entrez Gene | 18441501 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



