 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | evm_27.model.AmTr_v1.0_scaffold00029.323 | ||||||||
| Common Name | AMTR_s00029p00215750 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; basal Magnoliophyta; Amborellales; Amborellaceae; Amborella
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 286aa MW: 32475.3 Da PI: 9.5366 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 26.4 | 1.6e-08 | 15 | 60 | 4 | 47 |
S-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqky 47
WT eE +l+ +a +++ ++ +W+ Ia+ ++ g+t ++ ++ ++
evm_27.model.AmTr_v1.0_scaffold00029.323 15 WTLEENKLFEKALALYDKEtpdRWHMIAKIIP-GKTISDLMGHYKQL 60
*****************99*************.********999876 PP
| |||||||
| 2 | Myb_DNA-binding | 43.3 | 8.2e-14 | 118 | 162 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+W+++E+ l++ + k++G+g+W+ I+r + + Rt+ q+ s+ qky
evm_27.model.AmTr_v1.0_scaffold00029.323 118 PWSEDEHRLFLLGLKKHGKGDWRNISRNFVQSRTATQVASHAQKY 162
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 1.1E-7 | 11 | 63 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 9.33E-12 | 13 | 68 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 5.9E-4 | 14 | 62 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.4E-5 | 15 | 60 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.11E-7 | 15 | 61 | No hit | No description |
| PROSITE profile | PS50090 | 6.284 | 15 | 61 | IPR017877 | Myb-like domain |
| PROSITE profile | PS51294 | 19.381 | 111 | 167 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.3E-16 | 113 | 167 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.6E-16 | 114 | 165 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 7.5E-12 | 115 | 165 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-11 | 115 | 161 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 4.0E-11 | 118 | 162 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.06E-10 | 118 | 163 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 286 aa Download sequence Send to blast |
MEFHSSAPYF PSSIWTLEEN KLFEKALALY DKETPDRWHM IAKIIPGKTI SDLMGHYKQL 60 VDDISRIEAG WIELPSYNSS GFRLDWFSGD SQSVDGLKNG SRRASAKPPD HERKKGVPWS 120 EDEHRLFLLG LKKHGKGDWR NISRNFVQSR TATQVASHAQ KYFIRLSSGS KDKRRSSIHD 180 ITTANLSEKR SPSPPSPNIQ SQAFNTQLST SSDPIKTQNF NSFFSPNQTS EEVGSFISSV 240 PQAKATMYIS PPRTISPYEE KLHESDMGTQ HLMYRLQAYN LHNYC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 5e-16 | 15 | 81 | 11 | 77 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
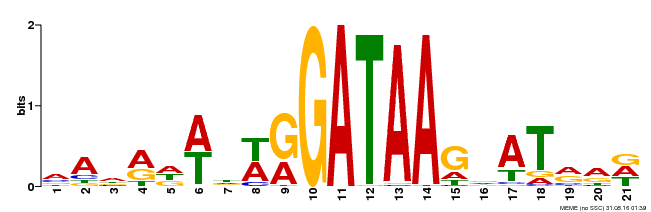
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006848111.2 | 0.0 | transcription factor DIVARICATA isoform X2 | ||||
| Refseq | XP_020525346.1 | 0.0 | transcription factor DIVARICATA isoform X1 | ||||
| Refseq | XP_020525347.1 | 0.0 | transcription factor DIVARICATA isoform X2 | ||||
| Swissprot | Q8S9H7 | 2e-77 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | W1PIA2 | 0.0 | W1PIA2_AMBTC; Uncharacterized protein | ||||
| STRING | ERN09692 | 0.0 | (Amborella trichopoda) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP275 | 17 | 122 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 3e-67 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | evm_27.model.AmTr_v1.0_scaffold00029.323 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



