 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G66300.1 | ||||||||
| Common Name | ANAC105, K1L20.8, NAC105, VND3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 292aa MW: 34320.5 Da PI: 6.4071 | ||||||||
| Description | NAC domain containing protein 105 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 173.1 | 8.5e-54 | 13 | 139 | 2 | 127 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyWkatgkdkevlskk 96
ppGfrFhPtdeelv +yLkkk+++++++l +vi+e+d+yk+ePwdL++ ++ e++ewyfFs+rdkky+tg+r+nrat +g+Wkatg+dk+v+ +
AT5G66300.1 13 PPGFRFHPTDEELVGYYLKKKIASQRIDL-DVIREIDLYKIEPWDLQErcRIGYeEQTEWYFFSHRDKKYPTGTRTNRATVAGFWKATGRDKAVYL-N 108
9****************************.9***************953433332667**************************************.9 PP
NAM 97 gelvglkktLvfykgrapkgektdWvmheyr 127
++l+g++ktLvfy+grap+g+k+dW++hey
AT5G66300.1 109 SKLIGMRKTLVFYRGRAPNGQKSDWIIHEYY 139
99***************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.07E-58 | 10 | 162 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 57.388 | 12 | 162 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.1E-26 | 13 | 138 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007275 | Biological Process | multicellular organism development | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:0071555 | Biological Process | cell wall organization | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000039 | anatomy | shoot axis vascular system | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0003011 | anatomy | root vascular system | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025275 | anatomy | procambium | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 292 aa Download sequence Send to blast |
MMKVDQDYSC SIPPGFRFHP TDEELVGYYL KKKIASQRID LDVIREIDLY KIEPWDLQER 60 CRIGYEEQTE WYFFSHRDKK YPTGTRTNRA TVAGFWKATG RDKAVYLNSK LIGMRKTLVF 120 YRGRAPNGQK SDWIIHEYYS LESHQNSPPQ EEGWVVCRAF KKRTTIPTKR RQLWDPNCLF 180 YDDATLLEPL DKRARHNPDF TATPFKQELL SEASHVQDGD FGSMYLQCID DDQFSQLPQL 240 ESPSLPSEIT PHSTTFSENS SRKDDMSSEK RITDWRYLDK FVASQFLMSG ED |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 5e-50 | 5 | 164 | 8 | 170 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 247084_at | 0.0 | ||||
| Expression Atlas | AT5G66300 | - | ||||
| AtGenExpress | AT5G66300 | - | ||||
| ATTED-II | AT5G66300 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Up-regulated during xylem vessel element formation. Expressed preferentially in procambial cells adjacent to root meristem. {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:25148240}. | |||||
| Uniprot | TISSUE SPECIFICITY: Detected in root protoxylem and metaxylem poles and in vessels of protoxylems, outermost metaxylems, inner metaxylems, shoots and hypocotyls. Expressed in roots, hypocotyls, cotyledons and leaves (PubMed:18445131). Present in developing xylems (PubMed:16103214, PubMed:17565617). Present in root developing xylems (PubMed:16103214). Specifically expressed in vessels but not in interfascicular fibers in stems (PubMed:25148240). {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:17565617, ECO:0000269|PubMed:18445131, ECO:0000269|PubMed:25148240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a NAC-domain transcription factor. Expressed in the vascular tissue. | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:25148240}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
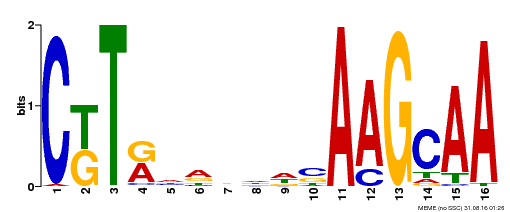
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00590 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G66300.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| IntAct | Search Q9FH59 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Reduced secondary wall thickening in vessels and collapsed vessel. {ECO:0000269|PubMed:25148240}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G66300 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT015824 | 0.0 | BT015824.1 Arabidopsis thaliana At5g66300 gene, complete cds. | |||
| GenBank | BT020216 | 0.0 | BT020216.1 Arabidopsis thaliana At5g66300 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_201431.1 | 0.0 | NAC domain containing protein 105 | ||||
| Swissprot | Q9FH59 | 0.0 | NC105_ARATH; NAC domain-containing protein 105 | ||||
| TrEMBL | A0A178UE15 | 0.0 | A0A178UE15_ARATH; VND3 | ||||
| STRING | AT5G66300.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM15134 | 16 | 19 | Representative plant | OGRP17 | 15 | 800 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G66300.1 |
| Entrez Gene | 836762 |
| iHOP | AT5G66300 |
| wikigenes | AT5G66300 |



