| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 64.8 | 9e-21 | 9 | 56 | 1 | 48 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklye 48
krienks rqvtfskRrng++ KA LS+LC+ +av+++s +gkly
AT5G65050.3 9 KRIENKSSRQVTFSKRRNGLIEKARQLSILCESSIAVLVVSGSGKLYK 56
79********************************************96 PP
|
| 2 | K-box | 39.3 | 2.8e-14 | 93 | 164 | 24 | 98 |
K-box 24 LkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkl 98
Lk+ +e q+ +l ++++ s+ L +Le+qLe++l+ R++K+el++ +++ lqk e+ l+een++L +++
AT5G65050.3 93 LKELLEIVQS---KLEESNVDNASVDTLISLEEQLETALSVTRARKTELMMGEVKSLQKTENLLREENQTLASQV 164
5555555554...47777899999***********************************************9987 PP
|
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
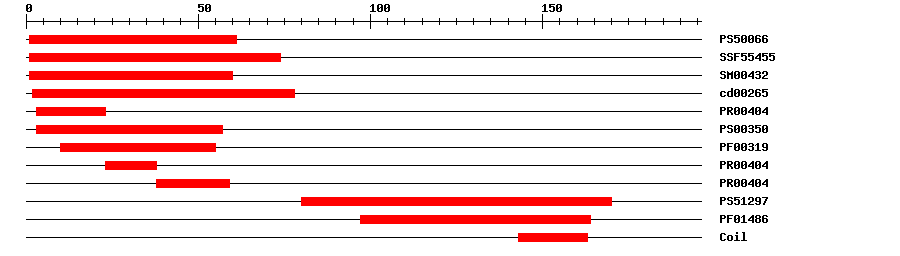
| PROSITE profile | PS50066 | 27.74 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 1.57E-25 | 1 | 74 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 7.7E-31 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.38E-31 | 2 | 78 | No hit | No description |
| PRINTS | PR00404 | 2.0E-23 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 8.7E-21 | 10 | 55 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.0E-23 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.0E-23 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 10.893 | 80 | 170 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 1.2E-10 | 97 | 164 | IPR002487 | Transcription factor, K-box |
| Publications
? help Back to Top |
- Alvarez-Buylla ER, et al.
MADS-box gene evolution beyond flowers: expression in pollen, endosperm, guard cells, roots and trichomes.
Plant J., 2000. 24(4): p. 457-66
[PMID:11115127] - Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Ratcliffe OJ,Kumimoto RW,Wong BJ,Riechmann JL
Analysis of the Arabidopsis MADS AFFECTING FLOWERING gene family: MAF2 prevents vernalization by short periods of cold.
Plant Cell, 2003. 15(5): p. 1159-69
[PMID:12724541] - Parenicová L, et al.
Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world.
Plant Cell, 2003. 15(7): p. 1538-51
[PMID:12837945] - Kofuji R, et al.
Evolution and divergence of the MADS-box gene family based on genome-wide expression analyses.
Mol. Biol. Evol., 2003. 20(12): p. 1963-77
[PMID:12949148] - Bae MS,Cho EJ,Choi EY,Park OK
Analysis of the Arabidopsis nuclear proteome and its response to cold stress.
Plant J., 2003. 36(5): p. 652-63
[PMID:14617066] - He Y,Doyle MR,Amasino RM
PAF1-complex-mediated histone methylation of FLOWERING LOCUS C chromatin is required for the vernalization-responsive, winter-annual habit in Arabidopsis.
Genes Dev., 2004. 18(22): p. 2774-84
[PMID:15520273] - Doyle MR, et al.
HUA2 is required for the expression of floral repressors in Arabidopsis thaliana.
Plant J., 2005. 41(3): p. 376-85
[PMID:15659097] - Kim SY, et al.
Establishment of the vernalization-responsive, winter-annual habit in Arabidopsis requires a putative histone H3 methyl transferase.
Plant Cell, 2005. 17(12): p. 3301-10
[PMID:16258034] - Andersson CR, et al.
The FLX gene of Arabidopsis is required for FRI-dependent activation of FLC expression.
Plant Cell Physiol., 2008. 49(2): p. 191-200
[PMID:18156133] - Alexandre CM,Hennig L
FLC or not FLC: the other side of vernalization.
J. Exp. Bot., 2008. 59(6): p. 1127-35
[PMID:18390846] - Caicedo AL,Richards C,Ehrenreich IM,Purugganan MD
Complex rearrangements lead to novel chimeric gene fusion polymorphisms at the Arabidopsis thaliana MAF2-5 flowering time gene cluster.
Mol. Biol. Evol., 2009. 26(3): p. 699-711
[PMID:19139056] - Rosloski SM,Jali SS,Balasubramanian S,Weigel D,Grbic V
Natural diversity in flowering responses of Arabidopsis thaliana caused by variation in a tandem gene array.
Genetics, 2010. 186(1): p. 263-76
[PMID:20551443] - Severing EI, et al.
Predicting the impact of alternative splicing on plant MADS domain protein function.
PLoS ONE, 2012. 7(1): p. e30524
[PMID:22295091] - Rosloski SM, et al.
Functional analysis of splice variant expression of MADS AFFECTING FLOWERING 2 of Arabidopsis thaliana.
Plant Mol. Biol., 2013. 81(1-2): p. 57-69
[PMID:23111501] - Suter L,Rüegg M,Zemp N,Hennig L,Widmer A
Gene regulatory variation mediates flowering responses to vernalization along an altitudinal gradient in Arabidopsis.
Plant Physiol., 2014. 166(4): p. 1928-42
[PMID:25339407] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Airoldi CA,McKay M,Davies B
MAF2 Is Regulated by Temperature-Dependent Splicing and Represses Flowering at Low Temperatures in Parallel with FLM.
PLoS ONE, 2015. 10(5): p. e0126516
[PMID:25955034]
|




